Assessing the Influence of Mutation on GTPase Transition States by Using X-ray Crystallography, (19) F NMR, and DFT Approaches.
Jin, Y., Molt, R.W., Pellegrini, E., Cliff, M.J., Bowler, M.W., Richards, N.G.J., Blackburn, G.M., Waltho, J.P.(2017) Angew Chem Int Ed Engl 56: 9732-9735
- PubMed: 28498638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.201703074
- Primary Citation Related Structures:
5M6X, 5M70 - PubMed Abstract:
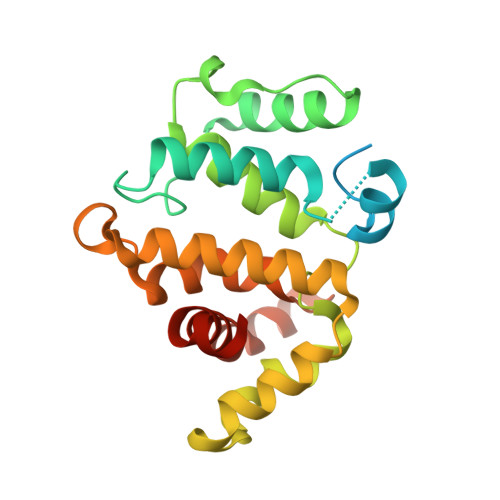
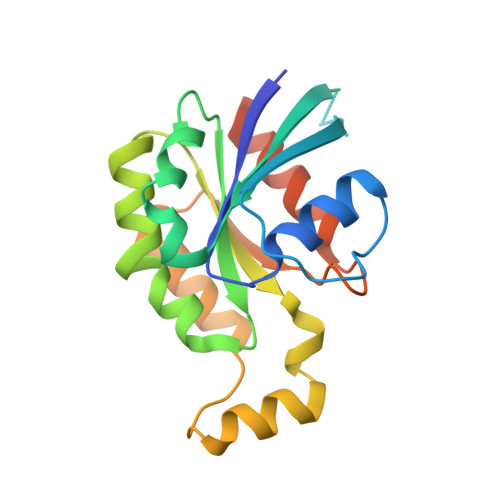
We report X-ray crystallographic and 19 F NMR studies of the G-protein RhoA complexed with MgF 3 - , GDP, and RhoGAP, which has the mutation Arg85'Ala. When combined with DFT calculations, these data permit the identification of changes in transition state (TS) properties. The X-ray data show how Tyr34 maintains solvent exclusion and the core H-bond network in the active site by relocating to replace the missing Arg85' sidechain. The 19 F NMR data show deshielding effects that indicate the main function of Arg85' is electronic polarization of the transferring phosphoryl group, primarily mediated by H-bonding to O 3G and thence to P G . DFT calculations identify electron-density redistribution and pinpoint why the TS for guanosine 5'-triphosphate (GTP) hydrolysis is higher in energy when RhoA is complexed with RhoGAP Arg85'Ala relative to wild-type (WT) RhoGAP. This study demonstrates that 19 F NMR measurements, in combination with X-ray crystallography and DFT calculations, can reliably dissect the response of small GTPases to site-specific modifications.
- Department of Molecular Biology and Biotechnology, Krebs Institute, University of Sheffield, Sheffield, S10 2TN, UK.
Organizational Affiliation: