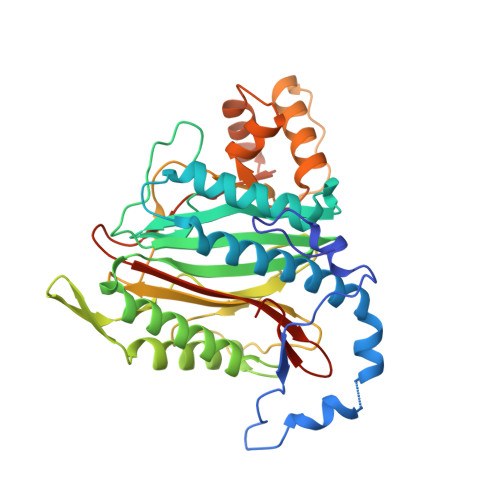
Novel reversible methionine aminopeptidase-2 (MetAP-2) inhibitors based on purine and related bicyclic templates.
Heinrich, T., Buchstaller, H.P., Cezanne, B., Rohdich, F., Bomke, J., Friese-Hamim, M., Krier, M., Knochel, T., Musil, D., Leuthner, B., Zenke, F.(2017) Bioorg Med Chem Lett 27: 551-556
- PubMed: 27998678 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.12.019
- Primary Citation Related Structures:
5LYW, 5LYX - PubMed Abstract:
The natural product fumagillin 1 and derivatives like TNP-470 2 or beloranib 3 bind to methionine aminopeptidase 2 (MetAP-2) irreversibly. This enzyme is critical for protein maturation and plays a key role in angiogenesis. In this paper we describe the synthesis, MetAP-2 binding affinity and structural analysis of reversible MetAP-2 inhibitors. Optimization of enzymatic activity of screening hit 10 (IC 50 : 1μM) led to the most potent compound 27 (IC 50 : 0.038μM), with a concomitant improvement in LLE from 2.1 to 4.2. Structural analysis of these MetAP-2 inhibitors revealed an unprecedented conformation of the His339 side-chain imidazole ring being co-planar sandwiched between the imidazole of His331 and the aryl-ether moiety, which is bound to the purine scaffold. Systematic alteration and reduction of H-bonding capability of this metal binding moiety induced an unexpected 180° flip for the triazolo[1,5-a]pyrimdine bicyclic template.
- Discovery Technologies, Merck KGaA, Frankfurter Str. 250, D-64293 Darmstadt, Germany. Electronic address: timo.heinrich@merckgroup.com.
Organizational Affiliation: