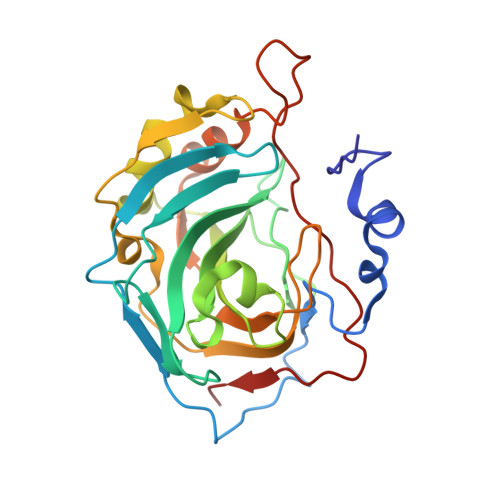
Self-Assembled Protein-Aromatic Foldamer Complexes with 2:3 and 2:2:1 Stoichiometries.
Jewginski, M., Granier, T., Langlois d'Estaintot, B., Fischer, L., Mackereth, C.D., Huc, I.(2017) J Am Chem Soc 139: 2928-2931
- PubMed: 28170240 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.7b00184
- Primary Citation Related Structures:
5L3O, 5L6K, 5LVS - PubMed Abstract:
The promotion of protein dimerization using the aggregation properties of a protein ligand was explored and shown to produce complexes with unusual stoichiometries. Helical foldamer 2 was synthesized and bound to human carbonic anhydrase (HCA) using a nanomolar active site ligand. Crystal structures show that the hydrophobicity of 2 and interactions of its side chains lead to the formation of an HCA 2 -2 3 complex in which three helices of 2 are stacked, two of them being linked to an HCA molecule. The middle foldamer in the stack can be replaced by alternate sequences 3 or 5. Solution studies by CD and NMR confirm left-handedness of the helical foldamers as well as HCA dimerization.
- CBMN (UMR5248), Univ. Bordeaux, CNRS, IPB , Institut Européen de Chimie et Biologie, 2 rue Robert Escarpit, 33600 Pessac, France.
Organizational Affiliation: