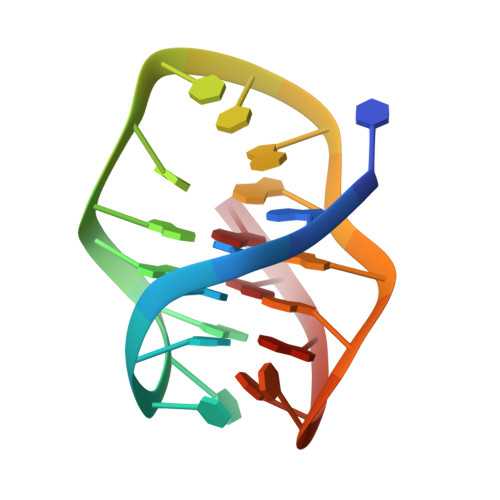
Reversible pH Switch of Two-Quartet G-Quadruplexes Formed by Human Telomere.
Galer, P., Wang, B., Sket, P., Plavec, J.(2016) Angew Chem Int Ed Engl 55: 1993-1997
- PubMed: 26836334 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201507569
- Primary Citation Related Structures:
5LQG, 5LQH - PubMed Abstract:
A four-repeat human telomere DNA sequence without the 3'-end guanine, d[TAGGG(TTAGGG)2 TTAGG] (htel1-ΔG23) has been found to adopt two distinct two G-quartet antiparallel basket-type G-quadruplexes, TD and KDH(+) in presence of KCl. NMR, CD, and UV spectroscopy have demonstrated that topology of KDH(+) form is distinctive with unique protonated T18⋅A20(+) ⋅G5 base triple and other capping structural elements that provide novel insight into structural polymorphism and heterogeneity of G-quadruplexes in general. Specific stacking interactions amongst two G-quartets flanking base triples and base pairs in TD and KDH(+) forms are reflected in 10 K higher thermal stability of KDH(+) . Populations of TD and KDH(+) forms are controlled by pH. The (de)protonation of A20 is the key for pH driven structural transformation of htel1-ΔG23. Reversibility offers possibilities for its utilization as a conformational switch within different compartments of living cell enabling specific ligand and protein interactions.
- Slovenian NMR Center, National Institute of Chemistry, Hajdrihova 19, Ljubljana, Slovenia.
Organizational Affiliation: