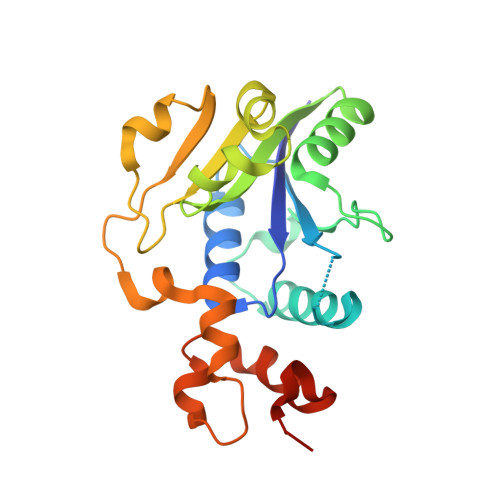
Structural and Functional Characterization of Plasmodium falciparum Nicotinic Acid Mononucleotide Adenylyltransferase.
Bathke, J., Fritz-Wolf, K., Brandstadter, C., Burkhardt, A., Jortzik, E., Rahlfs, S., Becker, K.(2016) J Mol Biology 428: 4946-4961
- PubMed: 27984041 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.10.023
- Primary Citation Related Structures:
5LLT, 5LM3 - PubMed Abstract:
Nicotinic acid mononucleotide adenylyltransferase (NaMNAT) is an indispensable enzyme for the synthesis of NAD and NAD phosphate. It catalyzes the adenylylation of nicotinic acid mononucleotide (NaMN) to yield nicotinic acid adenine dinucleotide (NaAD). Since NAD(H) and NAD phosphate(H) are essentially involved in metabolic and redox regulatory reactions, NaMNAT is an attractive drug target in the fight against bacterial and parasitic infections. Notably, NaMNAT of the malaria parasite Plasmodium falciparum possesses only 20% sequence identity with the homologous human enzyme. Here, we present for the first time the two X-ray structures of P. falciparum NaMNAT (PfNaMNAT)-in the product-bound state with NaAD and complexed with an α,β-non-hydrolizable ATP analog-the structures were determined to a resolution of 2.2Å and 2.5Å, respectively. The overall architecture of PfNaMNAT was found to be more similar to its bacterial homologs than its human counterparts although the PPHK motif conserved in bacteria is missing. Furthermore, PfNaMNAT possesses two cysteine residues within the active site that have not been described for any other NaMNATase so far and are likely to be involved in redox regulation of PfNaMNAT activity. Enzymatic studies and surface plasmon resonance data reveal that PfNaMNAT is capable of utilizing NaMN and nicotinamide mononucleotide with a slight preference for NaMN. Surprisingly, a comparison with the active site of Escherichia coli NaMNAT showed very similar architectures, despite different substrate preferences.
- Biochemistry and Molecular Biology, Interdisciplinary Research Center, Justus Liebig University, D-35392 Giessen, Germany.
Organizational Affiliation: