Discovery of a novel conformational equilibrium in urokinase-type plasminogen activator.
Kromann-Hansen, T., Louise Lange, E., Peter Srensen, H., Hassanzadeh-Ghassabeh, G., Huang, M., Jensen, J.K., Muyldermans, S., Declerck, P.J., Komives, E.A., Andreasen, P.A.(2017) Sci Rep 7: 3385-3385
- PubMed: 28611361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-03457-7
- Primary Citation Related Structures:
5LHN, 5LHP, 5LHQ, 5LHR, 5LHS - PubMed Abstract:
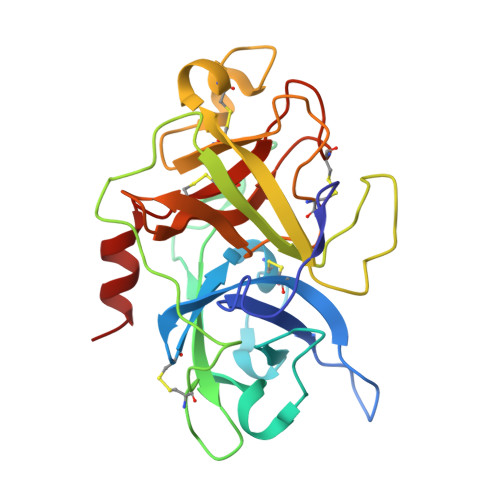
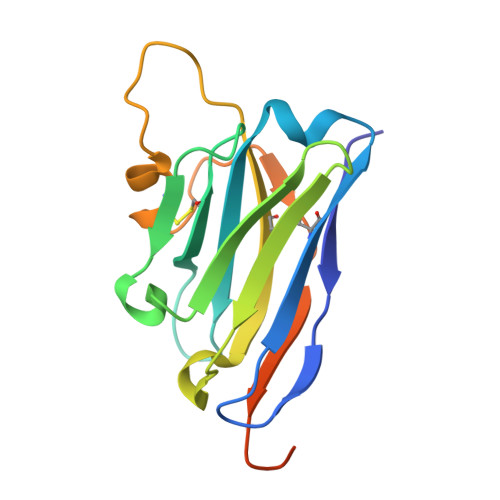
Although trypsin-like serine proteases have flexible surface-exposed loops and are known to adopt higher and lower activity conformations, structural determinants for the different conformations have remained largely obscure. The trypsin-like serine protease, urokinase-type plasminogen activator (uPA), is central in tissue remodeling processes and also strongly implicated in tumor metastasis. We solved five X-ray crystal structures of murine uPA (muPA) in the absence and presence of allosteric molecules and/or substrate-like molecules. The structure of unbound muPA revealed an unsuspected non-chymotrypsin-like protease conformation in which two β-strands in the core of the protease domain undergoes a major antiparallel-to-parallel conformational transition. We next isolated two anti-muPA nanobodies; an active-site binding nanobody and an allosteric nanobody. Crystal structures of the muPA:nanobody complexes and hydrogen-deuterium exchange mass spectrometry revealed molecular insights about molecular factors controlling the antiparallel-to-parallel equilibrium in muPA. Together with muPA activity assays, the data provide valuable insights into regulatory mechanisms and conformational flexibility of uPA and trypsin-like serine proteases in general.
- From the Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, California, United States. tkromanntofting@ucsd.edu.
Organizational Affiliation: