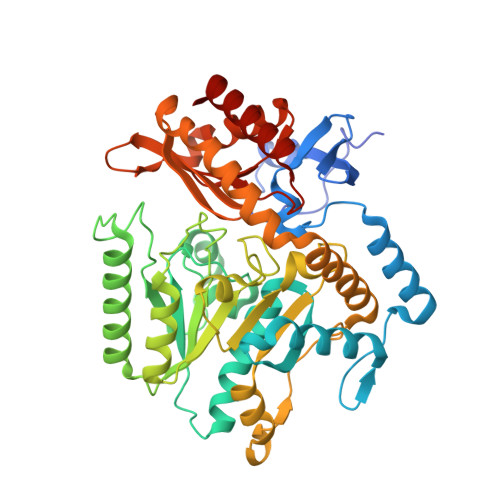
Explaining Operational Instability of Amine Transaminases: Substrate-Induced Inactivation Mechanism and Influence of Quaternary Structure on Enzyme-Cofactor Intermediate Stability.
Borner, T., Ramisch, S., Reddem, E., Bartsch, S., Vogel, A., Thunnissen, A.M.W.H., Adlercreutz, P., Grey, C.(2017) ACS Catal 7: 1259-1269