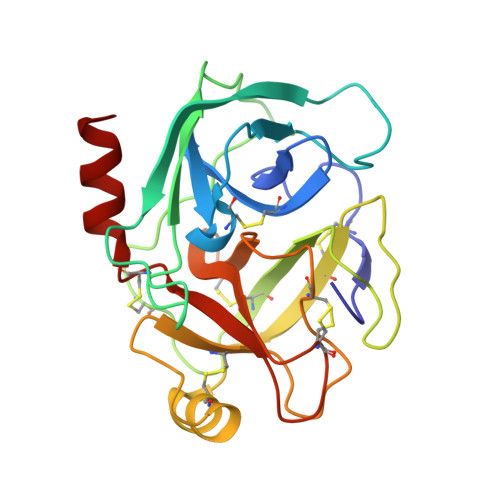
Trypsin inhibitors for the treatment of pancreatitis.
Brandl, T., Simic, O., Skaanderup, P.R., Namoto, K., Berst, F., Ehrhardt, C., Schiering, N., Mueller, I., Woelcke, J.(2016) Bioorg Med Chem Lett 26: 4340-4344
- PubMed: 27476144 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.07.029
- Primary Citation Related Structures:
5LGO, 5LH4, 5LH8 - PubMed Abstract:
Proline-based trypsin inhibitors occupying the S1-S2-S1' region were identified by an HTS screening campaign. It was discovered that truncation of the P1' moiety and appropriate extension into the S4 region led to highly potent trypsin inhibitors with excellent selectivity against related serine proteases and a favorable hERG profile.
- Novartis Pharma AG, Novartis Institutes for BioMedical Research, Novartis Campus, CH-4002 Basel, Switzerland. Electronic address: trixi.brandl@novartis.com.
Organizational Affiliation: