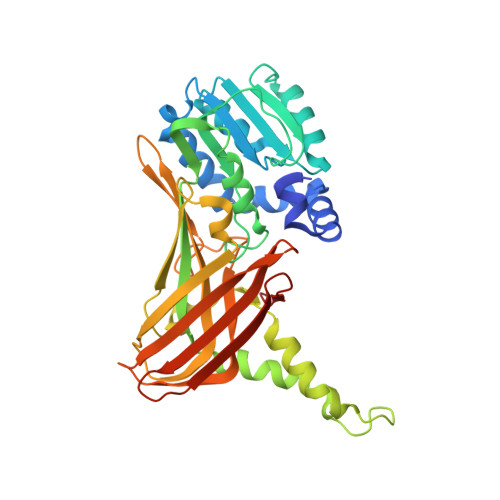
Transition state mimics are valuable mechanistic probes for structural studies with the arginine methyltransferase CARM1.
van Haren, M.J., Marechal, N., Troffer-Charlier, N., Cianciulli, A., Sbardella, G., Cavarelli, J., Martin, N.I.(2017) Proc Natl Acad Sci U S A 114: 3625-3630
- PubMed: 28330993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1618401114
- Primary Citation Related Structures:
5LGP, 5LGQ, 5LGR, 5LGS - PubMed Abstract:
Coactivator associated arginine methyltransferase 1 (CARM1) is a member of the protein arginine methyltransferase (PRMT) family and methylates a range of proteins in eukaryotic cells. Overexpression of CARM1 is implicated in a number of cancers, and it is therefore seen as a potential therapeutic target. Peptide sequences derived from the well-defined CARM1 substrate poly(A)-binding protein 1 (PABP1) were covalently linked to an adenosine moiety as in the AdoMet cofactor to generate transition state mimics. These constructs were found to be potent CARM1 inhibitors and also formed stable complexes with the enzyme. High-resolution crystal structures of CARM1 in complex with these compounds confirm a mode of binding that is indeed reflective of the transition state at the CARM1 active site. Given the transient nature of PRMT-substrate complexes, such transition state mimics represent valuable chemical tools for structural studies aimed at deciphering the regulation of arginine methylation mediated by the family of arginine methyltransferases.
- Department of Chemical Biology & Drug Discovery, Utrecht Institute for Pharmaceutical Sciences, Utrecht University, 3584 CG Utrecht, The Netherlands.
Organizational Affiliation: