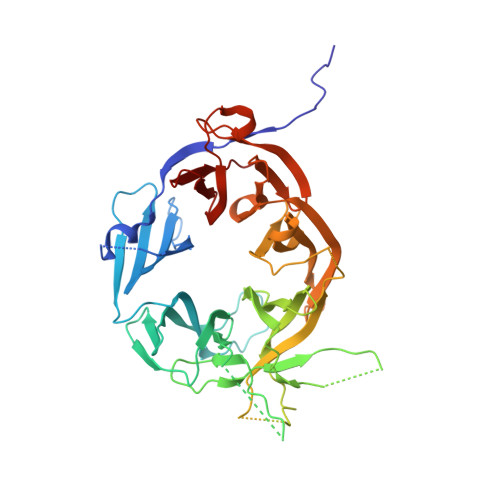
WD40 domain of Apc1 is critical for the coactivator-induced allosteric transition that stimulates APC/C catalytic activity.
Li, Q., Chang, L., Aibara, S., Yang, J., Zhang, Z., Barford, D.(2016) Proc Natl Acad Sci U S A 113: 10547-10552
- PubMed: 27601667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1607147113
- Primary Citation Related Structures:
5LGG - PubMed Abstract:
The anaphase-promoting complex/cyclosome (APC/C) is a large multimeric cullin-RING E3 ubiquitin ligase that orchestrates cell-cycle progression by targeting cell-cycle regulatory proteins for destruction via the ubiquitin proteasome system. The APC/C assembly comprises two scaffolding subcomplexes: the platform and the TPR lobe that together coordinate the juxtaposition of the catalytic and substrate-recognition modules. The platform comprises APC/C subunits Apc1, Apc4, Apc5, and Apc15. Although the role of Apc1 as an APC/C scaffolding subunit has been characterized, its specific functions in contributing toward APC/C catalytic activity are not fully understood. Here, we report the crystal structure of the N-terminal domain of human Apc1 (Apc1N) determined at 2.2-Å resolution and provide an atomic-resolution description of the architecture of its WD40 (WD40 repeat) domain (Apc1(WD40)). To understand how Apc1(WD40) contributes to APC/C activity, a mutant form of the APC/C with Apc1(WD40) deleted was generated and evaluated biochemically and structurally. We found that the deletion of Apc1(WD40) abolished the UbcH10-dependent ubiquitination of APC/C substrates without impairing the Ube2S-dependent ubiquitin chain elongation activity. A cryo-EM structure of an APC/C-Cdh1 complex with Apc1(WD40) deleted showed that the mutant APC/C is locked into an inactive conformation in which the UbcH10-binding site of the catalytic module is inaccessible. Additionally, an EM density for Apc15 is not visible. Our data show that Apc1(WD40) is required to mediate the coactivator-induced conformational change of the APC/C that is responsible for stimulating APC/C catalytic activity by promoting UbcH10 binding. In contrast, Ube2S activity toward APC/C substrates is not dependent on the initiation-competent conformation of the APC/C.
- Medical Research Council Laboratory of Molecular Biology, Cambridge CB2 0QH, United Kingdom; Section of Structural Biology, Chester Beatty Laboratories, Institute of Cancer Research, London SW3 6JB, United Kingdom.
Organizational Affiliation: