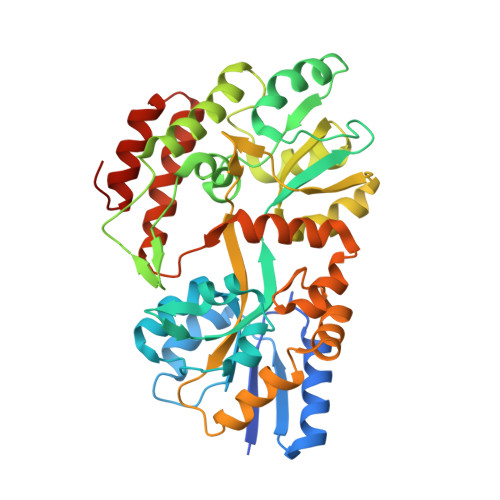
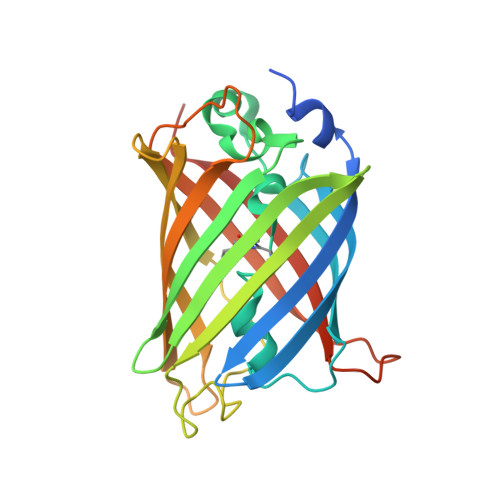
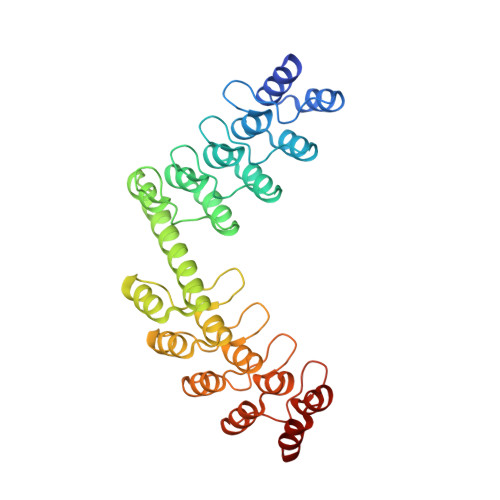Rigidly connected multispecific artificial binders with adjustable geometries.
Wu, Y., Batyuk, A., Honegger, A., Brandl, F., Mittl, P.R.E., Pluckthun, A.(2017) Sci Rep 7: 11217-11217
- PubMed: 28894181 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-11472-x
- Primary Citation Related Structures:
5LE2, 5LE3, 5LE4, 5LE6, 5LE7, 5LE8, 5LE9, 5LEA, 5LEB, 5LEC, 5LED, 5LEE, 5LEL, 5LEM, 5LW2 - PubMed Abstract:
Multivalent binding proteins can gain biological activities beyond what is inherent in the individual binders, by bringing together different target molecules, restricting their conformational flexibility or changing their subcellular localization. In this study, we demonstrate a method to build up rigid multivalent and multispecific scaffolds by exploiting the modular nature of a repeat protein scaffold and avoiding flexible linkers. We use DARPins (Designed Ankyrin Repeat Proteins), synthetic binding proteins based on the Ankyrin-repeat protein scaffold, as binding units. Their ease of in vitro selection, high production yield and stability make them ideal specificity-conferring building blocks for the design of more complex constructs. C- and N-terminal DARPin capping repeats were re-designed to be joined by a shared helix in such a way that rigid connector modules are formed. This allows us to join two or more DARPins in predefined geometries without compromising their binding affinities and specificities. Nine connector modules with distinct geometries were designed; for eight of these we were able to confirm the structure by X-ray crystallography, while only one did not crystallize. The bispecific constructs were all able to bind both target proteins simultaneously.
- Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, CH-8057, Zürich, Switzerland.
Organizational Affiliation: