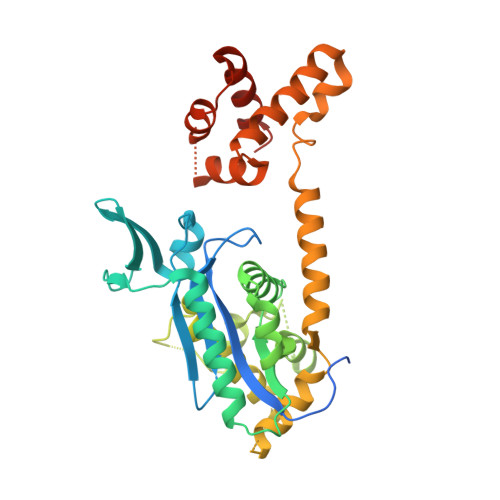
The bicoid mRNA localization factor Exuperantia is an RNA-binding pseudonuclease.
Lazzaretti, D., Veith, K., Kramer, K., Basquin, C., Urlaub, H., Irion, U., Bono, F.(2016) Nat Struct Mol Biol 23: 705-713
- PubMed: 27376588 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3254
- Primary Citation Related Structures:
5L7Z, 5L80 - PubMed Abstract:
Anterior patterning in Drosophila is mediated by the localization of bicoid (bcd) mRNA at the anterior pole of the oocyte. Exuperantia (Exu) is a putative exonuclease (EXO) associated with bcd and required for its localization. We present the crystal structure of Exu, which reveals a dimeric assembly with each monomer consisting of a 3'-5' EXO-like domain and a sterile alpha motif (SAM)-like domain. The catalytic site is degenerate and inactive. Instead, the EXO-like domain mediates dimerization and RNA binding. We show that Exu binds RNA directly in vitro, that the SAM-like domain is required for RNA binding activity and that Exu binds a structured element present in the bcd 3' untranslated region with high affinity. Through structure-guided mutagenesis, we show that Exu dimerization is essential for bcd localization. Our data demonstrate that Exu is a noncanonical RNA-binding protein with EXO-SAM-like domain architecture that interacts with its target RNA as a homodimer.
- Max Planck Institute for Developmental Biology, Tübingen, Germany.
Organizational Affiliation: