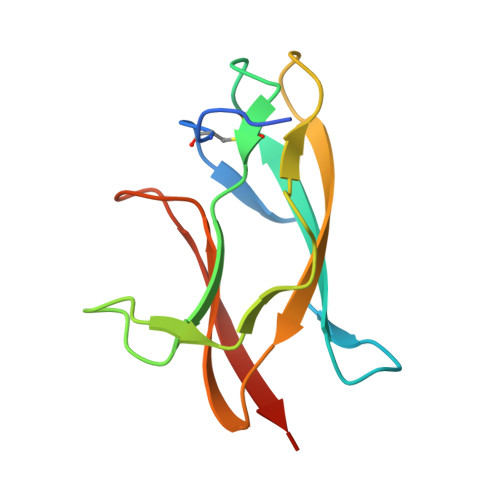
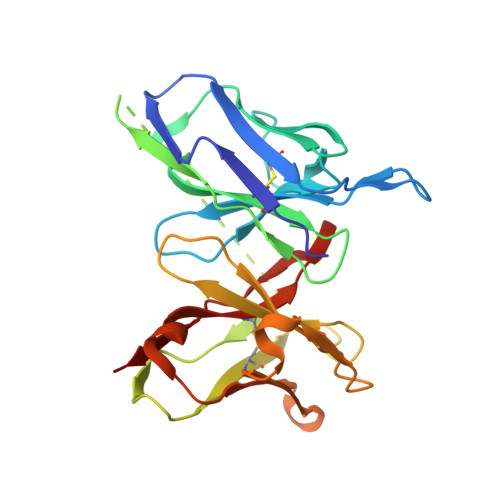
Structural Basis of Zika Virus-Specific Antibody Protection.
Zhao, H., Fernandez, E., Dowd, K.A., Speer, S.D., Platt, D.J., Gorman, M.J., Govero, J., Nelson, C.A., Pierson, T.C., Diamond, M.S., Fremont, D.H.(2016) Cell 166: 1016-1027
- PubMed: 27475895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2016.07.020
- Primary Citation Related Structures:
5KVD, 5KVE, 5KVF, 5KVG - PubMed Abstract:
Zika virus (ZIKV) infection during pregnancy has emerged as a global public health problem because of its ability to cause severe congenital disease. Here, we developed six mouse monoclonal antibodies (mAbs) against ZIKV including four (ZV-48, ZV-54, ZV-64, and ZV-67) that were ZIKV specific and neutralized infection of African, Asian, and American strains to varying degrees. X-ray crystallographic and competition binding analyses of Fab fragments and scFvs defined three spatially distinct epitopes in DIII of the envelope protein corresponding to the lateral ridge (ZV-54 and ZV-67), C-C' loop (ZV-48 and ZV-64), and ABDE sheet (ZV-2) regions. In vivo passive transfer studies revealed protective activity of DIII-lateral ridge specific neutralizing mAbs in a mouse model of ZIKV infection. Our results suggest that DIII is targeted by multiple type-specific antibodies with distinct neutralizing activity, which provides a path for developing prophylactic antibodies for use in pregnancy or designing epitope-specific vaccines against ZIKV.
- Department of Pathology & Immunology, Washington University School of Medicine, Saint Louis, MO 63110, USA.
Organizational Affiliation: