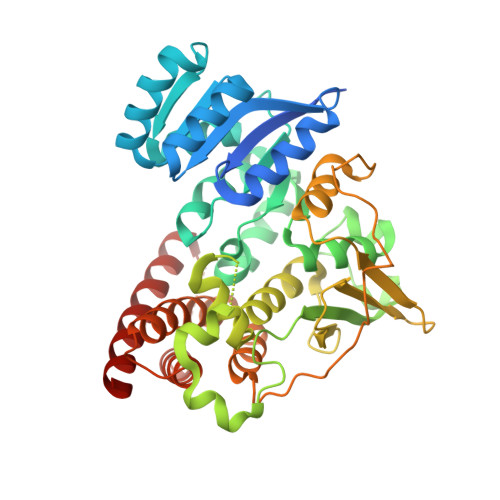
Structural characterization of 1-deoxy-D-xylulose 5-phosphate Reductoisomerase from Vibrio vulnificus.
Ussin, N.K., Bagnell, A.M., Offermann, L.R., Abdulsalam, R., Perdue, M.L., Magee, P., Chruszcz, M.(2018) Biochim Biophys Acta Proteins Proteom 1866: 1209-1215
- PubMed: 30278288 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2018.09.008
- Primary Citation Related Structures:
5KQO, 5KRR, 5KRV, 5KRY, 5KS1 - PubMed Abstract:
Vibrio vulnificus, a gram-negative bacterium, is the leading cause of seafood-borne illnesses and mortality in the United States. Previous studies have identified metabolites 2-C-methylerythritol 4-phosphate (MEP) as being essential for V. vulnificus growth and function. It was shown that 1-deoxy-D-xylulose-5-phosphate reductoisomerase (Dxr) is a critical enzyme in the viability of V. vulnificus, and many other bacteria, as it catalyzes the rearrangement of 1-deoxy-D-xylulose-5-phosphate (Dxp) to 2-C-methylerythritol 4-phosphate (MEP) within the MEP pathway, found in plants and bacteria. The MEP pathway produces the isoprenoids, isopentenyl diphosphate and dimethylallyl pyrophosphate. In this study, we produced and structurally characterized V. vulnificus Dxr. The enzyme forms a dimeric assembly and contains a metal ion in the active site. Protein produced in Escherichia coli co-purifies with Mg 2+ ions, however the Mg 2+ cations may be substituted with Mn 2+ , as both of these metals may be utilized by Dxrs. These findings will provide a basis for the design of Dxr inhibitors that may find application as antimicrobial compounds.
- Department of Chemistry and Biochemistry, University of South Carolina, Columbia, SC 29208, United States.
Organizational Affiliation: