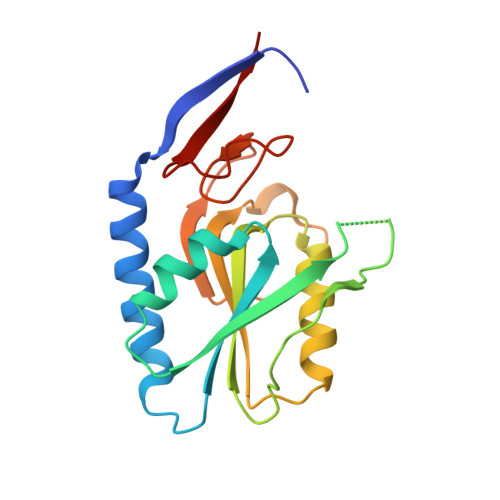
Crystal Structures of Acyclic Nucleoside Phosphonates in Complex with Escherichia coli Hypoxanthine Phosphoribosyltransferase
Eng, W.S., Hockova, D., Spacek, P., Baszczynski, O., Janeba, Z., Naesens, L., Keough, D.T., Guddat, L.W.(2016) ChemistrySelect 1: 6267-6276