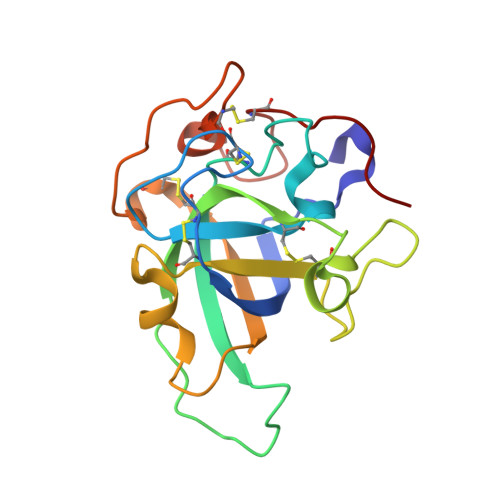
Structure, computational and biochemical analysis of PcCel45A endoglucanase from Phanerochaete chrysosporium and catalytic mechanisms of GH45 subfamily C members.
Godoy, A.S., Pereira, C.S., Ramia, M.P., Silveira, R.L., Camilo, C.M., Kadowaki, M.A., Lange, L., Busk, P.K., Nascimento, A.S., Skaf, M.S., Polikarpov, I.(2018) Sci Rep 8: 3678-3678
- PubMed: 29487297 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-21798-9
- Primary Citation Related Structures:
5KJQ - PubMed Abstract:
The glycoside hydrolase family 45 (GH45) of carbohydrate modifying enzymes is mostly comprised of β-1,4-endoglucanases. Significant diversity between the GH45 members has prompted the division of this family into three subfamilies: A, B and C, which may differ in terms of the mechanism, general architecture, substrate binding and cleavage. Here, we use a combination of X-ray crystallography, bioinformatics, enzymatic assays, molecular dynamics simulations and site-directed mutagenesis experiments to characterize the structure, substrate binding and enzymatic specificity of the GH45 subfamily C endoglucanase from Phanerochaete chrysosporium (PcCel45A). We investigated the role played by different residues in the binding of the enzyme to cellulose oligomers of different lengths and examined the structural characteristics and dynamics of PcCel45A that make subfamily C so dissimilar to other members of the GH45 family. Due to the structural similarity shared between PcCel45A and domain I of expansins, comparative analysis of their substrate binding was also carried out. Our bioinformatics sequence analyses revealed that the hydrolysis mechanisms in GH45 subfamily C is not restricted to use of the imidic asparagine as a general base in the "Newton's cradle" catalytic mechanism recently proposed for this subfamily.
- São Carlos Institute of Physics, University of São Paulo, São Carlos 13566-590, São Paulo, Brazil.
Organizational Affiliation: