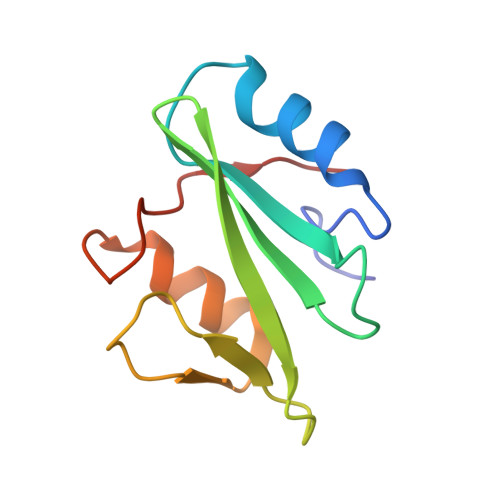
The X-ray Crystallographic Structure of Human EAT2 (SH2D1B).
Taha, M., Nezerwa, E., Nam, H.J.(2016) Protein Pept Lett 23: 862-866
- PubMed: 27586300 Search on PubMed
- DOI: https://doi.org/10.2174/0929866523666160831162239
- Primary Citation Related Structures:
5KAZ - PubMed Abstract:
Ewing's Sarcoma transcript-2 (EAT2) also known as SH2D1B is involved in regulation of signalling lymphocytic activation molecule (SLAM) family receptor functions. Cytoplasmic tails of SLAM family receptors contain tyrosine residues which mediate the downstream signal transduction through their phosphorylation. EAT2, composed of a single SH2 domain and a short C-terminal tail, binds to the phosphotyrosine residues and regulates SLAM family receptor signalling. We have determined the crystal structure of the human EAT2 protein in an unliganded form. Compared with the mouse EAT2-peptide complex structure, we observe conformational differences in the loops involved in ligand binding. When compared with SAP, the other single SH2 domain protein in human, EAT2 shows similar binding energies to unphosphorylated ligands. This is inconsistent to the previous data showing low affinity of EAT2 toward unphosphorylated peptides compared to SAP which shows high affinity. Additional factors other than the SH2 domains may contribute to the reported differences.
- Department of Bioengineering, The University of Texas at Dallas, 800 W. Campbell Rd, RL10, Richardson, USA. hnam@utdallas.edu.
Organizational Affiliation: