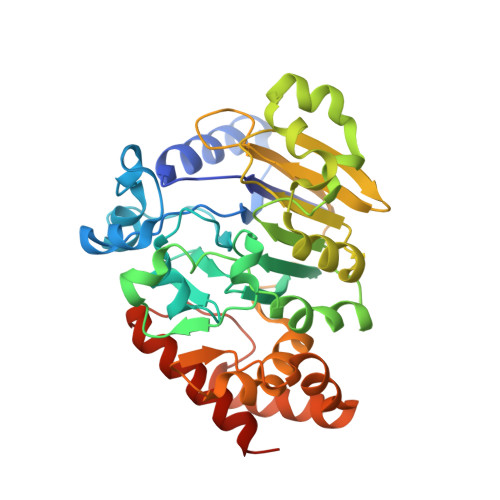
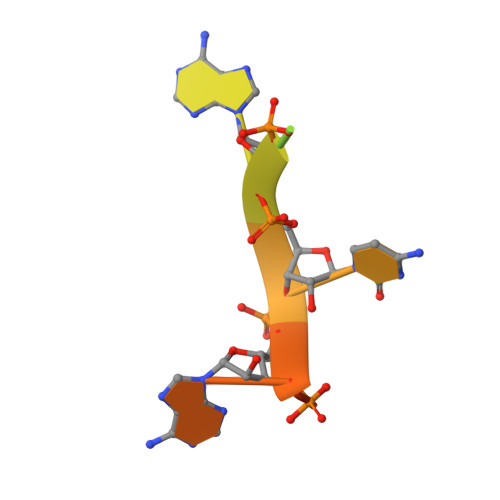
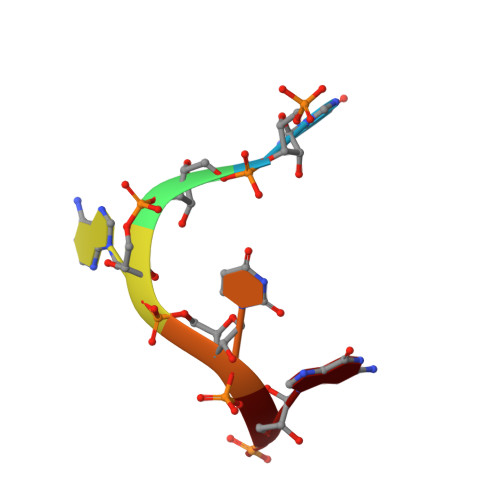
The RNA lariat debranching enzyme Dbr1: metal dependence and branched RNA co-crystal structures
Clark, N.E., Katolik, A., Roberts, K., Taylor, A.B., Holloway, S.P., Schuermann, J.P., Montemayor, E.J., Stevens, S.W., Fitzpatrick, P.F., Damha, M.J., Hart, P.J.(2016) Proc Natl Acad Sci U S A