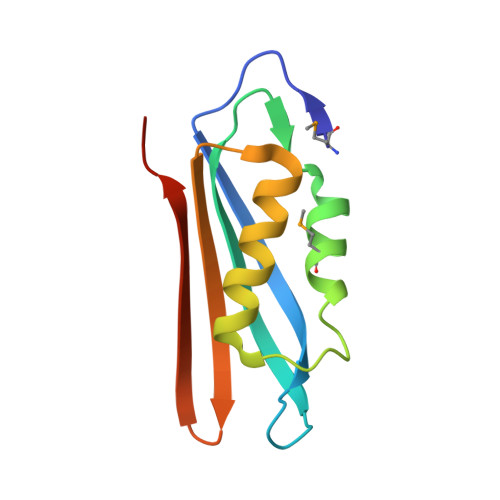
Crystal structure of the archaeosine synthase QueF-like-Insights into amidino transfer and tRNA recognition by the tunnel fold.
Mei, X., Alvarez, J., Bon Ramos, A., Samanta, U., Iwata-Reuyl, D., Swairjo, M.A.(2017) Proteins 85: 103-116
- PubMed: 27802572 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.25202
- Primary Citation Related Structures:
5JYX, 5K0P - PubMed Abstract:
The tunneling-fold (T-fold) structural superfamily has emerged as a versatile protein scaffold of diverse catalytic activities. This is especially evident in the pathways to the 7-deazaguanosine modified nucleosides of tRNA queuosine and archaeosine. Four members of the T-fold superfamily have been confirmed in these pathways and here we report the crystal structure of a fifth enzyme; the recently discovered amidinotransferase QueF-Like (QueF-L), responsible for the final step in the biosynthesis of archaeosine in the D-loop of tRNA in a subset of Crenarchaeota. QueF-L catalyzes the conversion of the nitrile group of the 7-cyano-7-deazaguanine (preQ 0 ) base of preQ 0 -modified tRNA to a formamidino group. The structure, determined in the presence of preQ 0 , reveals a symmetric T-fold homodecamer of two head-to-head facing pentameric subunits, with 10 active sites at the inter-monomer interfaces. Bound preQ 0 forms a stable covalent thioimide bond with a conserved active site cysteine similar to the intermediate previously observed in the nitrile reductase QueF. Despite distinct catalytic functions, phylogenetic distributions, and only 19% sequence identity, the two enzymes share a common preQ 0 binding pocket, and likely a common mechanism of thioimide formation. However, due to tight twisting of its decamer, QueF-L lacks the NADPH binding site present in QueF. A large positively charged molecular surface and a docking model suggest simultaneous binding of multiple tRNA molecules and structure-specific recognition of the D-loop by a surface groove. The structure sheds light on the mechanism of nitrile amidation, and the evolution of diverse chemistries in a common fold. Proteins 2016; 85:103-116. © 2016 Wiley Periodicals, Inc.
- Department of Chemistry and Biochemistry, San Diego State University- 5500 Campanile Drive, San Diego, California, 92182.
Organizational Affiliation: