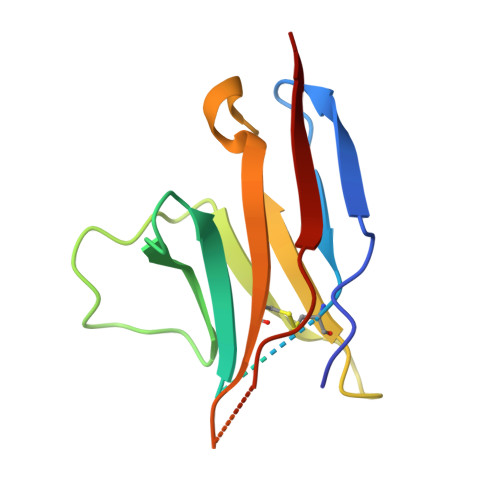
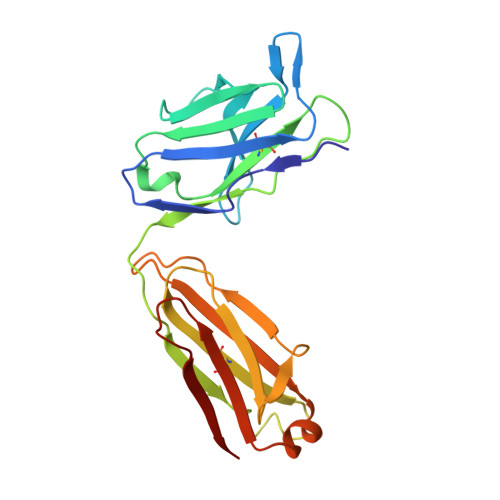
Structural basis for blocking PD-1-mediated immune suppression by therapeutic antibody pembrolizumab.
Na, Z., Yeo, S.P., Bharath, S.R., Bowler, M.W., Balijkcij, E., Wang, C.I., Song, H.(2017) Cell Res 27: 147-150
- PubMed: 27325296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2016.77
- Primary Citation Related Structures:
5JXE - Institute of Molecular and Cell Biology, 61 Biopolis Drive, Singapore 138673, Singapore.
Organizational Affiliation: