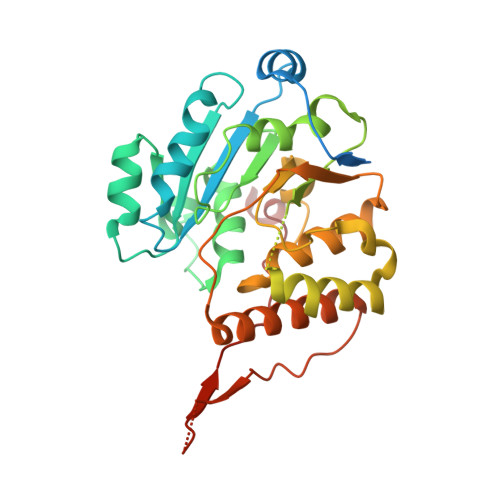
Crystal structure of glucosyl-3-phosphoglycerate synthase from Mycobacterium tuberculosis - apo form
Albesa-Jove, D., Romero-Garcia, J., Sancho-Vaello, E., Contreras, F.-X., Rodrigo-Unzueta, A., Comino, N., Carreras-Gonzalez, A., Arrasate, P., Urresti, S., Biarnes, X., Planas, A., Guerin, M.E.To be published.