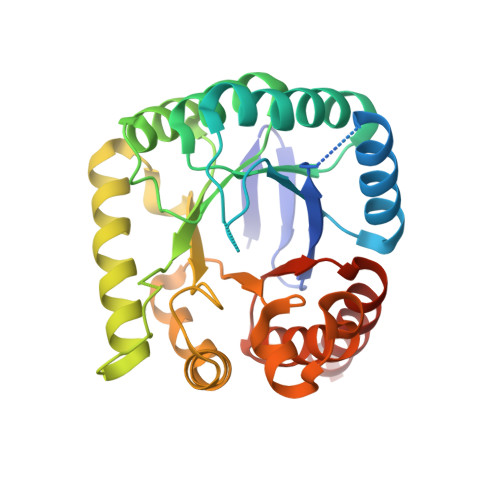
Pterin-sulfa conjugates as dihydropteroate synthase inhibitors and antibacterial agents.
Zhao, Y., Shadrick, W.R., Wallace, M.J., Wu, Y., Griffith, E.C., Qi, J., Yun, M.K., White, S.W., Lee, R.E.(2016) Bioorg Med Chem Lett 26: 3950-3954
- PubMed: 27423480 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2016.07.006
- Primary Citation Related Structures:
5JQ9 - PubMed Abstract:
The sulfonamide class of antibiotics has been in continuous use for over 70years. They are thought to act by directly inhibiting dihydropteroate synthase (DHPS), and also acting as prodrugs that sequester pterin pools by forming dead end pterin-sulfonamide conjugates. In this study, eight pterin-sulfonamide conjugates were synthesized using a novel synthetic strategy and their biochemical and microbiological properties were investigated. The conjugates were shown to competitively inhibit DHPS, and inhibition was enhanced by the presence of pyrophosphate that is crucial to catalysis and is known to promote an ordering of the DHPS active site. The co-crystal structure of Yersinia pestis DHPS bound to one of the more potent conjugates revealed a mode of binding that is similar to that of the enzymatic product analog pteroic acid. The antimicrobial activities of the pterin-sulfonamide conjugates were measured against Escherichia coli in the presence and absence of folate precursors and dependent metabolites. These results show that the conjugates have appreciable antibacterial activity and act by an on target, anti-folate pathway mechanism rather than as simple dead end products.
- Department of Chemical Biology and Therapeutics, St. Jude Children's Research Hospital, 262 Danny Thomas Place, Mail Stop 1000, Memphis, TN 38105, United States.
Organizational Affiliation: