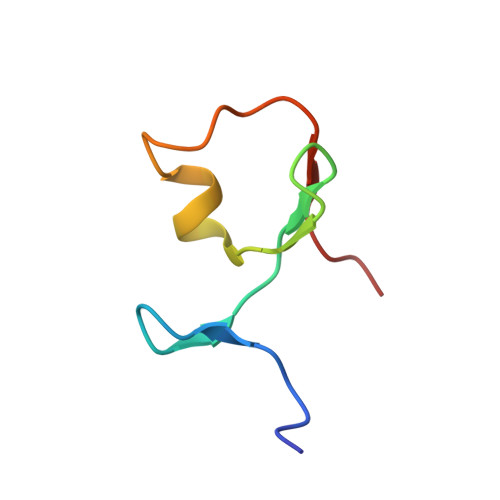
Solution NMR structure of the TRIM21 B-box2 and identification of residues involved in its interaction with the RING domain.
Wallenhammar, A., Anandapadamanaban, M., Lemak, A., Mirabello, C., Lundstrom, P., Wallner, B., Sunnerhagen, M.(2017) PLoS One 12: e0181551-e0181551
- PubMed: 28753623 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0181551
- Primary Citation Related Structures:
5JPX - PubMed Abstract:
Tripartite motif-containing (TRIM) proteins are defined by the sequential arrangement of RING, B-box and coiled-coil domains (RBCC), where the B-box domain is a unique feature of the TRIM protein family. TRIM21 is an E3 ubiquitin-protein ligase implicated in innate immune signaling by acting as an autoantigen and by modifying interferon regulatory factors. Here we report the three-dimensional solution structure of the TRIM21 B-box2 domain by nuclear magnetic resonance (NMR) spectroscopy. The structure of the B-box2 domain, comprising TRIM21 residues 86-130, consists of a short α-helical segment with an N-terminal short β-strand and two anti-parallel β-strands jointly found the core, and adopts a RING-like fold. This ββαβ core largely defines the overall fold of the TRIM21 B-box2 and the coordination of one Zn2+ ion stabilizes the tertiary structure of the protein. Using NMR titration experiments, we have identified an exposed interaction surface, a novel interaction patch where the B-box2 is likely to bind the N-terminal RING domain. Our structure together with comparisons with other TRIM B-box domains jointly reveal how its different surfaces are employed for various modular interactions, and provides extended understanding of how this domain relates to flanking domains in TRIM proteins.
- Division of Chemistry, Department of Physics, Chemistry and Biology, Linköping University, Linköping, Sweden.
Organizational Affiliation: