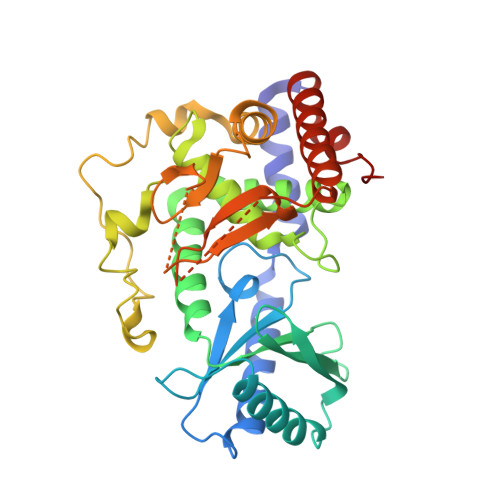
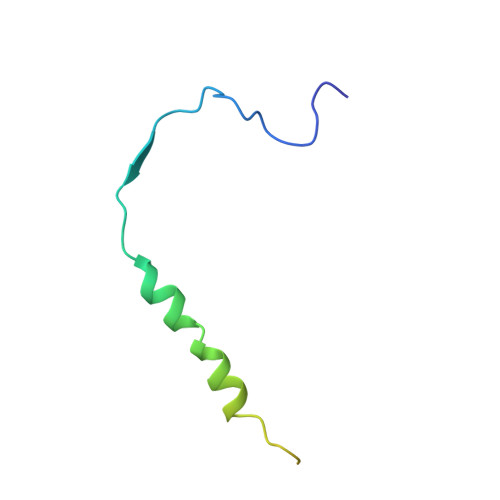
Structural basis for the antagonistic roles of RNP-8 and GLD-3 in GLD-2 poly(A)-polymerase activity.
Nakel, K., Bonneau, F., Basquin, C., Habermann, B., Eckmann, C.R., Conti, E.(2016) RNA 22: 1139-1145
- PubMed: 27288313 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.056598.116
- Primary Citation Related Structures:
5JNB - PubMed Abstract:
Cytoplasmic polyadenylation drives the translational activation of specific mRNAs in early metazoan development and is performed by distinct complexes that share the same catalytic poly(A)-polymerase subunit, GLD-2. The activity and specificity of GLD-2 depend on its binding partners. In Caenorhabditis elegans, GLD-2 promotes spermatogenesis when bound to GLD-3 and oogenesis when bound to RNP-8. GLD-3 and RNP-8 antagonize each other and compete for GLD-2 binding. Following up on our previous mechanistic studies of GLD-2-GLD-3, we report here the 2.5 Å resolution structure and biochemical characterization of a GLD-2-RNP-8 core complex. In the structure, RNP-8 embraces the poly(A)-polymerase, docking onto several conserved hydrophobic hotspots present on the GLD-2 surface. RNP-8 stabilizes GLD-2 and indirectly stimulates polyadenylation. RNP-8 has a different amino-acid sequence and structure as compared to GLD-3. Yet, it binds the same surfaces of GLD-2 by forming alternative interactions, rationalizing the remarkable versatility of GLD-2 complexes.
- Department of Structural Cell Biology, Max-Planck-Institute of Biochemistry, D-82152 Martinsried, Germany.
Organizational Affiliation: