Discovery of a Chemical Probe Bisamide (CCT251236): An Orally Bioavailable Efficacious Pirin Ligand from a Heat Shock Transcription Factor 1 (HSF1) Phenotypic Screen.
Cheeseman, M.D., Chessum, N.E., Rye, C.S., Pasqua, A.E., Tucker, M.J., Wilding, B., Evans, L.E., Lepri, S., Richards, M., Sharp, S.Y., Ali, S., Rowlands, M., O'Fee, L., Miah, A., Hayes, A., Henley, A.T., Powers, M., Te Poele, R., De Billy, E., Pellegrino, L., Raynaud, F., Burke, R., van Montfort, R.L., Eccles, S.A., Workman, P., Jones, K.(2017) J Med Chem 60: 180-201
- PubMed: 28004573 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01055
- Primary Citation Related Structures:
5JCT - PubMed Abstract:
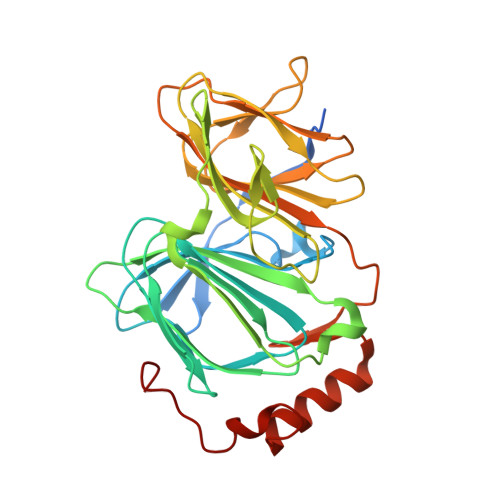
Phenotypic screens, which focus on measuring and quantifying discrete cellular changes rather than affinity for individual recombinant proteins, have recently attracted renewed interest as an efficient strategy for drug discovery. In this article, we describe the discovery of a new chemical probe, bisamide (CCT251236), identified using an unbiased phenotypic screen to detect inhibitors of the HSF1 stress pathway. The chemical probe is orally bioavailable and displays efficacy in a human ovarian carcinoma xenograft model. By developing cell-based SAR and using chemical proteomics, we identified pirin as a high affinity molecular target, which was confirmed by SPR and crystallography.
- Cancer Research UK Cancer Therapeutics Unit at The Institute of Cancer Research , London SW7 3RP, United Kingdom.
Organizational Affiliation: