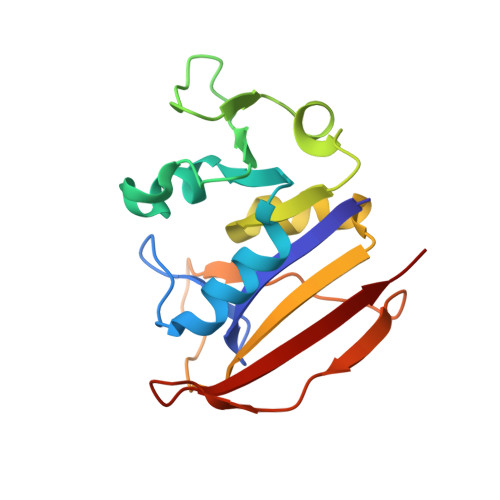
Propargyl-Linked Antifolates Are Potent Inhibitors of Drug-Sensitive and Drug-Resistant Mycobacterium tuberculosis.
Hajian, B., Scocchera, E., Keshipeddy, S., G-Dayanandan, N., Shoen, C., Krucinska, J., Reeve, S., Cynamon, M., Anderson, A.C., Wright, D.L.(2016) PLoS One 11: e0161740-e0161740
- PubMed: 27580226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0161740
- Primary Citation Related Structures:
5JA3 - PubMed Abstract:
Mycobacterium tuberculosis continues to cause widespread, life-threatening disease. In the last decade, this threat has grown dramatically as multi- and extensively-drug resistant (MDR and XDR) bacteria have spread globally and the number of agents that effectively treat these infections is significantly reduced. We have been developing the propargyl-linked antifolates (PLAs) as potent inhibitors of the essential enzyme dihydrofolate reductase (DHFR) from bacteria and recently found that charged PLAs with partial zwitterionic character showed improved mycobacterial cell permeability. Building on a hypothesis that these PLAs may penetrate the outer membrane of M. tuberculosis and inhibit the essential cytoplasmic DHFR, we screened a group of PLAs for antitubercular activity. In this work, we identified several PLAs as potent inhibitors of the growth of M. tuberculosis with several of the compounds exhibiting minimum inhibition concentrations equal to or less than 1 μg/mL. Furthermore, two of the compounds were very potent inhibitors of MDR and XDR strains. A high resolution crystal structure of one PLA bound to DHFR from M. tuberculosis reveals the interactions of the ligands with the target enzyme.
- Department of Pharmaceutical Sciences, University of Connecticut, Storrs, Connecticut, United States of America.
Organizational Affiliation: