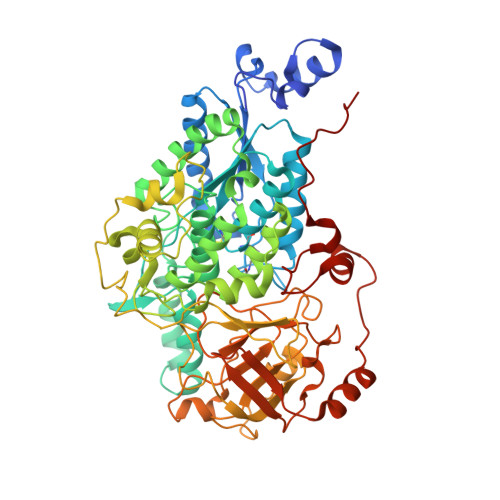
The Crystal Structure of a Bacterial l-Arabinonate Dehydratase Contains a [2Fe-2S] Cluster.
Rahman, M.M., Andberg, M., Thangaraj, S.K., Parkkinen, T., Penttila, M., Janis, J., Koivula, A., Rouvinen, J., Hakulinen, N.(2017) ACS Chem Biol 12: 1919-1927
- PubMed: 28574691 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.7b00304
- Primary Citation Related Structures:
5J83, 5J84, 5J85 - PubMed Abstract:
We present a novel crystal structure of the IlvD/EDD family enzyme, l-arabinonate dehydratase from Rhizobium leguminosarum bv. trifolii (RlArDHT, EC 4.2.1.25), which catalyzes the conversion of l-arabinonate to 2-dehydro-3-deoxy-l-arabinonate. The enzyme is a tetramer consisting of a dimer of dimers, where each monomer is composed of two domains. The active site contains a catalytically important [2Fe-2S] cluster and Mg 2+ ion and is buried between two domains, and also at the dimer interface. The active site Lys129 was found to be carbamylated. Ser480 and Thr482 were shown to be essential residues for catalysis, and the S480A mutant structure showed an unexpected open conformation in which the active site was more accessible for the substrate. This structure showed the partial binding of l-arabinonate, which allowed us to suggest that the alkoxide ion form of the Ser480 side chain functions as a base and the [2Fe-2S] cluster functions as a Lewis acid in the elimination reaction.
- Department of Chemistry, University of Eastern Finland , P.O. Box 111, FIN-80101 Joensuu, Finland.
Organizational Affiliation: