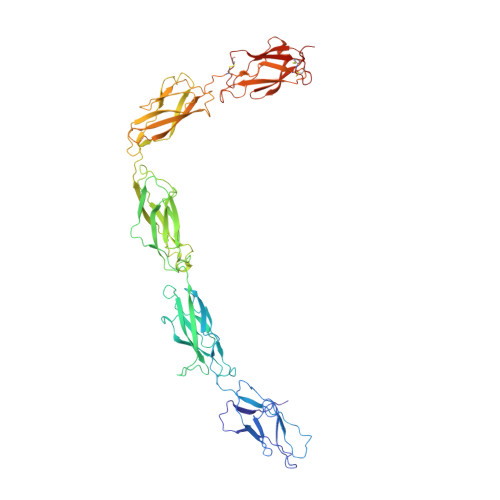
Structural basis of adhesive binding by desmocollins and desmogleins.
Harrison, O.J., Brasch, J., Lasso, G., Katsamba, P.S., Ahlsen, G., Honig, B., Shapiro, L.(2016) Proc Natl Acad Sci U S A 113: 7160-7165
- PubMed: 27298358 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1606272113
- Primary Citation Related Structures:
5EQX, 5ERD, 5ERP, 5IRY, 5J5J - PubMed Abstract:
Desmosomes are intercellular adhesive junctions that impart strength to vertebrate tissues. Their dense, ordered intercellular attachments are formed by desmogleins (Dsgs) and desmocollins (Dscs), but the nature of trans-cellular interactions between these specialized cadherins is unclear. Here, using solution biophysics and coated-bead aggregation experiments, we demonstrate family-wise heterophilic specificity: All Dsgs form adhesive dimers with all Dscs, with affinities characteristic of each Dsg:Dsc pair. Crystal structures of ectodomains from Dsg2 and Dsg3 and from Dsc1 and Dsc2 show binding through a strand-swap mechanism similar to that of homophilic classical cadherins. However, conserved charged amino acids inhibit Dsg:Dsg and Dsc:Dsc interactions by same-charge repulsion and promote heterophilic Dsg:Dsc interactions through opposite-charge attraction. These findings show that Dsg:Dsc heterodimers represent the fundamental adhesive unit of desmosomes and provide a structural framework for understanding desmosome assembly.
- Department of Systems Biology, Columbia University, New York, NY 10032; Howard Hughes Medical Institute, Columbia University, New York, NY 10032;
Organizational Affiliation: