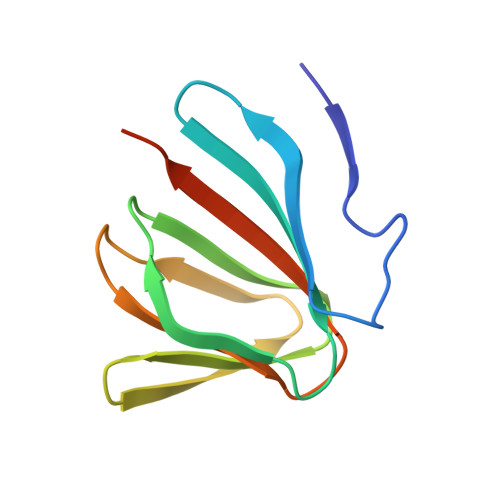
Structural identification of the lipopolysaccharide-binding capability of a cupin-family protein from Helicobacter pylori
Sim, D.W., Kim, J.H., Kim, H.Y., Jang, J.H., Lee, W.C., Kim, E.H., Park, P.J., Lee, K.H., Won, H.S.(2016) FEBS Lett 590: 2997-3004
- PubMed: 27466800 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12332
- Primary Citation Related Structures:
5J4F, 5J4G - PubMed Abstract:
We solved the crystal structure of a functionally uncharacterized protein, HP0902, from Helicobacter pylori. Its structure demonstrated an all-β cupin fold that cannot bind metal ions due to the absence of a metal-binding histidine that is conserved in many metallo-cupins. In contrast, isothermal titration calorimetry and NMR titration demonstrated that HP0902 is able to bind bacterial endotoxin lipopolysaccharides (LPS) through its surface-exposed loops, where metal-binding sites are usually found in other metallo-cupins. This report constitutes the first identification of an LPS-interacting protein, both in the cupin family and in H. pylori. Furthermore, identification of the ability of HP0902 to bind LPS uncovers a putative role for this protein in H. pylori pathogenicity.
- Department of Biotechnology, Research Institute (RIBHS) and College of Biomedical and Health Science, Konkuk University, Chungbuk, Korea.
Organizational Affiliation: