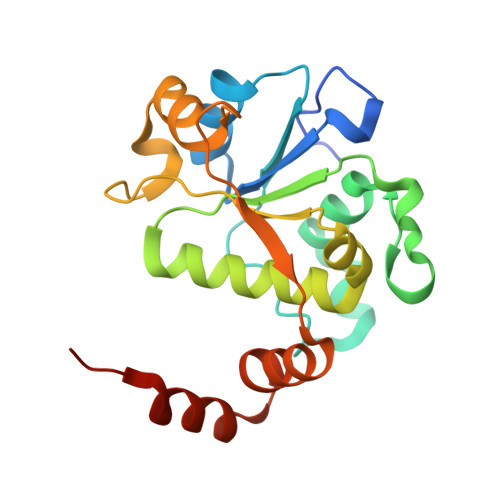
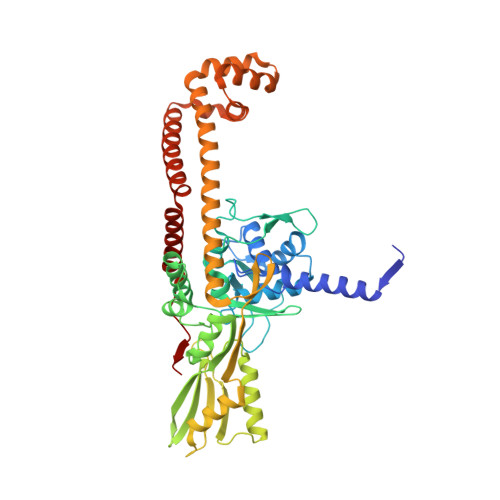
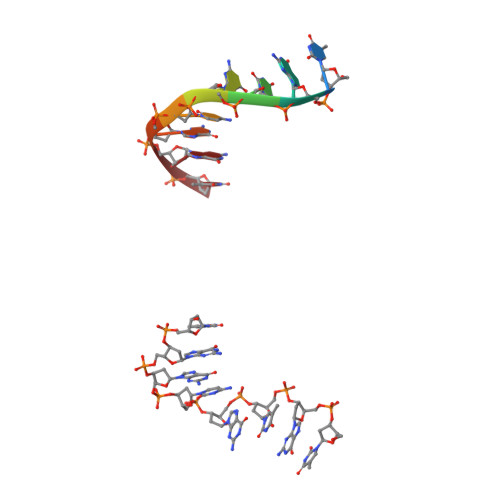
Novel tricyclics (e.g., GSK945237) as potent inhibitors of bacterial type IIA topoisomerases.
Miles, T.J., Hennessy, A.J., Bax, B., Brooks, G., Brown, B.S., Brown, P., Cailleau, N., Chen, D., Dabbs, S., Davies, D.T., Esken, J.M., Giordano, I., Hoover, J.L., Jones, G.E., Kusalakumari Sukmar, S.K., Markwell, R.E., Minthorn, E.A., Rittenhouse, S., Gwynn, M.N., Pearson, N.D.(2016) Bioorg Med Chem Lett 26: 2464-2469
- PubMed: 27055939 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2016.03.106
- Primary Citation Related Structures:
5IWI, 5IWM - PubMed Abstract:
During the course of our research on the lead optimisation of the NBTI (Novel Bacterial Type II Topoisomerase Inhibitors) class of antibacterials, we discovered a series of tricyclic compounds that showed good Gram-positive and Gram-negative potency. Herein we will discuss the various subunits that were investigated in this series and report advanced studies on compound 1 (GSK945237) which demonstrates good PK and in vivo efficacy properties.
- Diseases of the Developing World CEDD, GlaxoSmithKline, Calle Severo Ochoa, 2, 28760 Tres Cantos, Madrid, Spain. Electronic address: tim.j.miles@gsk.com.
Organizational Affiliation: