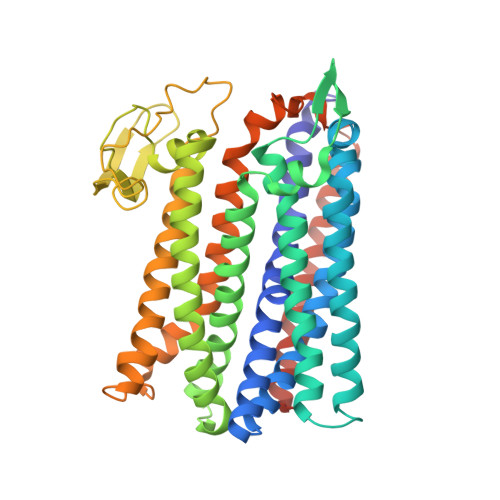
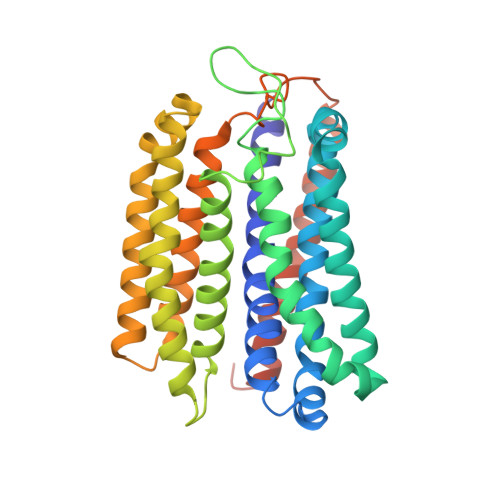
Structure of a bd oxidase indicates similar mechanisms for membrane-integrated oxygen reductases.
Safarian, S., Rajendran, C., Muller, H., Preu, J., Langer, J.D., Ovchinnikov, S., Hirose, T., Kusumoto, T., Sakamoto, J., Michel, H.(2016) Science 352: 583-586
- PubMed: 27126043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aaf2477
- Primary Citation Related Structures:
5DOQ, 5IR6 - PubMed Abstract:
The cytochrome bd oxidases are terminal oxidases that are present in bacteria and archaea. They reduce molecular oxygen (dioxygen) to water, avoiding the production of reactive oxygen species. In addition to their contribution to the proton motive force, they mediate viability under oxygen-related stress conditions and confer tolerance to nitric oxide, thus contributing to the virulence of pathogenic bacteria. Here we present the atomic structure of the bd oxidase from Geobacillus thermodenitrificans, revealing a pseudosymmetrical subunit fold. The arrangement and order of the heme cofactors support the conclusions from spectroscopic measurements that the cleavage of the dioxygen bond may be mechanistically similar to that in the heme-copper-containing oxidases, even though the structures are completely different.
- Department of Molecular Membrane Biology, Max Planck Institute of Biophysics, Max-von-Laue-Straße 3, D-60438 Frankfurt/Main, Germany.
Organizational Affiliation: