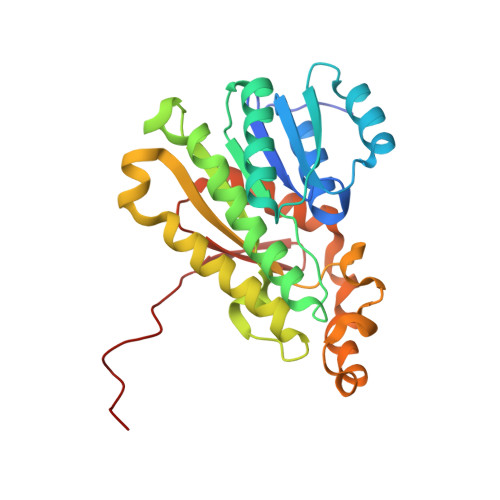
Structural Insights into the Drosophila melanogaster Retinol Dehydrogenase, a Member of the Short-Chain Dehydrogenase/Reductase Family.
Hofmann, L., Tsybovsky, Y., Alexander, N.S., Babino, D., Leung, N.Y., Montell, C., Banerjee, S., von Lintig, J., Palczewski, K.(2016) Biochemistry 55: 6545-6557
- PubMed: 27809489 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00907
- Primary Citation Related Structures:
5ILG, 5ILO - PubMed Abstract:
The 11-cis-retinylidene chromophore of visual pigments isomerizes upon interaction with a photon, initiating a downstream cascade of signaling events that ultimately lead to visual perception. 11-cis-Retinylidene is regenerated through enzymatic transformations collectively called the visual cycle. The first and rate-limiting enzymatic reaction within this cycle, i.e., the reduction of all-trans-retinal to all-trans-retinol, is catalyzed by retinol dehydrogenases. Here, we determined the structure of Drosophila melanogaster photoreceptor retinol dehydrogenase (PDH) isoform C that belongs to the short-chain dehydrogenase/reductase (SDR) family. This is the first reported structure of a SDR that possesses this biologically important activity. Two crystal structures of the same enzyme grown under different conditions revealed a novel conformational change of the NAD + cofactor, likely representing a change during catalysis. Amide hydrogen-deuterium exchange of PDH demonstrated changes in the structure of the enzyme upon dinucleotide binding. In D. melanogaster, loss of PDH activity leads to photoreceptor degeneration that can be partially rescued by transgenic expression of human RDH12. Based on the structure of PDH, we analyzed mutations causing Leber congenital amaurosis 13 in a homology model of human RDH12 to obtain insights into the molecular basis of RDH12 disease-causing mutations.
- Department of Pharmacology and Cleveland Center for Membrane and Structural Biology, School of Medicine, Case Western Reserve University , 10900 Euclid Avenue, Cleveland, Ohio 44106, United States.
Organizational Affiliation: