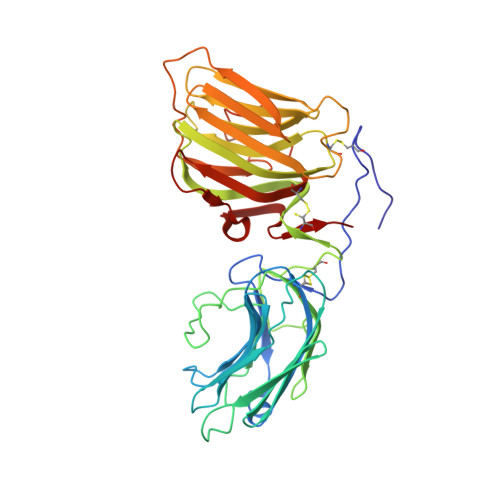
Structural basis of laminin binding to the LARGE glycans on dystroglycan.
Briggs, D.C., Yoshida-Moriguchi, T., Zheng, T., Venzke, D., Anderson, M.E., Strazzulli, A., Moracci, M., Yu, L., Hohenester, E., Campbell, K.P.(2016) Nat Chem Biol 12: 810-814
- PubMed: 27526028 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio.2146
- Primary Citation Related Structures:
5IK4, 5IK5, 5IK7, 5IK8 - PubMed Abstract:
Dystroglycan is a highly glycosylated extracellular matrix receptor with essential functions in skeletal muscle and the nervous system. Reduced matrix binding by α-dystroglycan (α-DG) due to perturbed glycosylation is a pathological feature of several forms of muscular dystrophy. Like-acetylglucosaminyltransferase (LARGE) synthesizes the matrix-binding heteropolysaccharide [-glucuronic acid-β1,3-xylose-α1,3-]n. Using a dual exoglycosidase digestion, we confirm that this polysaccharide is present on native α-DG from skeletal muscle. The atomic details of matrix binding were revealed by a high-resolution crystal structure of laminin-G-like (LG) domains 4 and 5 (LG4 and LG5) of laminin-α2 bound to a LARGE-synthesized oligosaccharide. A single glucuronic acid-β1,3-xylose disaccharide repeat straddles a Ca(2+) ion in the LG4 domain, with oxygen atoms from both sugars replacing Ca(2+)-bound water molecules. The chelating binding mode accounts for the high affinity of this protein-carbohydrate interaction. These results reveal a previously uncharacterized mechanism of carbohydrate recognition and provide a structural framework for elucidating the mechanisms underlying muscular dystrophy.
- Department of Life Sciences, Imperial College London, London, UK.
Organizational Affiliation: