Platinum(ii) O,S complexes as potential metallodrugs against Cisplatin resistance.
Hildebrandt, J., Hafner, N., Gorls, H., Kritsch, D., Ferraro, G., Durst, M., Runnebaum, I.B., Merlino, A., Weigand, W.(2016) Dalton Trans 45: 18876-18891
- PubMed: 27897281 Search on PubMed
- DOI: https://doi.org/10.1039/c6dt01388k
- Primary Citation Related Structures:
5IHG, 5II3, 5ILC, 5ILF - PubMed Abstract:
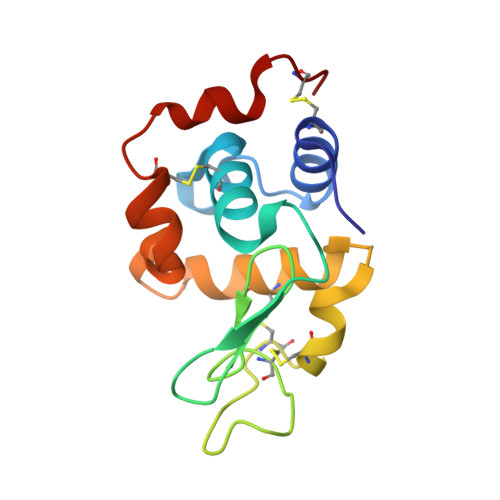
We report on platinum(ii) complexes with different cinnamic acid derivatives as ligands with cytotoxic activity against Cisplatin resistant ovarian cancer cell line subcultures of SKOV3 and A2780. A typical mechanism of action for platinum(ii) complexes as Cisplatin itself is binding to the DNA and inducing double-strand breaks. We examined the biological behavior of these potential drugs with 9-methylguanine using NMR spectroscopic methods and their DNA damage potential including γH2AX-foci analyses. X-ray diffraction methods have been used to elucidate the molecular structures of the platinum(ii) complexes. Interactions with the model protein lysozyme have been evaluated by different techniques including UV-Vis absorption spectroscopy, fluorescence and X-ray crystallography.
- Institut für Anorganische und Analytische Chemie Friedrich-Schiller-Universität Jena Humboldtstraße 8, 07743 Jena, Germany. wolfgang.weigand@uni-jena.de.
Organizational Affiliation: