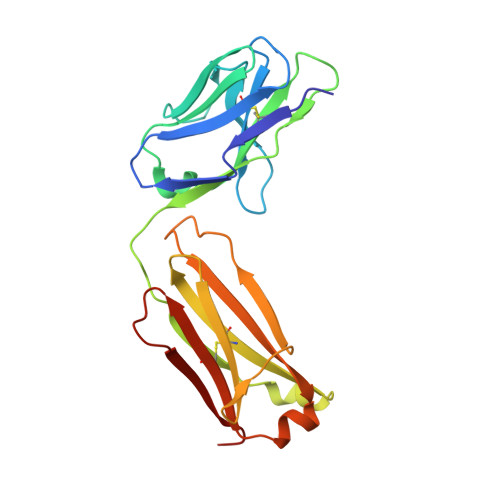
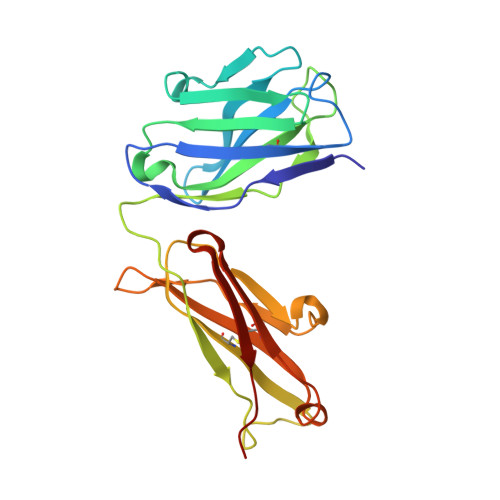
Bacterial production and structure-functional validation of a recombinant antigen-binding fragment (Fab) of an anti-cancer therapeutic antibody targeting epidermal growth factor receptor.
Kim, J.H., Sim, D.W., Park, D., Jung, T.G., Lee, S., Oh, T., Ha, J.R., Seok, S.H., Seo, M.D., Kang, H.C., Kim, Y.P., Won, H.S.(2016) Appl Microbiol Biotechnol 100: 10521-10529
- PubMed: 27470143 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-016-7717-z
- Primary Citation Related Structures:
5I76 - PubMed Abstract:
Fragment engineering of monoclonal antibodies (mAbs) has emerged as an excellent paradigm to develop highly efficient therapeutic and/or diagnostic agents. Engineered mAb fragments can be economically produced in bacterial systems using recombinant DNA technologies. In this work, we established recombinant production in Escherichia coli for monovalent antigen-binding fragment (Fab) adopted from a clinically used anticancer mAB drug cetuximab targeting epidermal growth factor receptor (EGFR). Recombinant DNA constructs were designed to express both polypeptide chains comprising Fab in a single vector and to secrete them to bacterial periplasmic space for efficient folding. Particularly, a C-terminal engineering to confer an interchain disulfide bond appeared to be able to enhance its heterodimeric integrity and EGFR-binding activity. Conformational relevance of the purified final product was validated by mass spectrometry and crystal structure at 1.9 Å resolution. Finally, our recombinant cetuximab-Fab was found to have strong binding affinity to EGFR overexpressed in human squamous carcinoma model (A431) cells. Its binding ability was comparable to that of cetuximab. Its EGFR-binding affinity was estimated at approximately 0.7 nM of Kd in vitro, which was quite stronger than the binding affinity of natural ligand EGF. Hence, the results validate that our construction could serve as an efficient platform to produce a recombinant cetuximab-Fab with a retained antigen-binding functionality.
- Department of Biotechnology, Research Institute (RIBHS) and College of Biomedical and Health Science, Konkuk University, Chungju, Chungbuk, 27478, South Korea.
Organizational Affiliation: