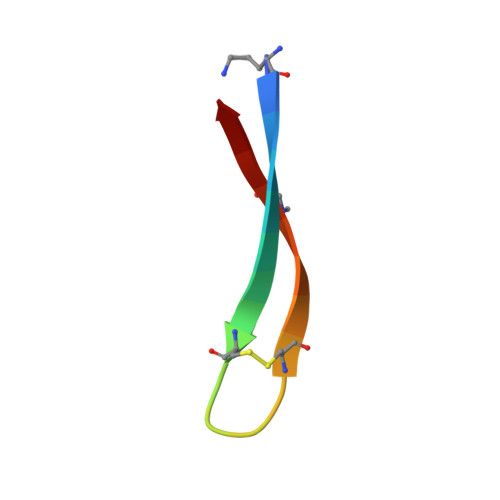
X-ray Crystallographic Structures of a Trimer, Dodecamer, and Annular Pore Formed by an A beta 17-36 beta-Hairpin.
Kreutzer, A.G., Hamza, I.L., Spencer, R.K., Nowick, J.S.(2016) J Am Chem Soc 138: 4634-4642
- PubMed: 26967810 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.6b01332
- Primary Citation Related Structures:
5HOW, 5HOX, 5HOY - PubMed Abstract:
High-resolution structures of oligomers formed by the β-amyloid peptide Aβ are needed to understand the molecular basis of Alzheimer's disease and develop therapies. This paper presents the X-ray crystallographic structures of oligomers formed by a 20-residue peptide segment derived from Aβ. The development of a peptide in which Aβ17-36 is stabilized as a β-hairpin is described, and the X-ray crystallographic structures of oligomers it forms are reported. Two covalent constraints act in tandem to stabilize the Aβ17-36 peptide in a hairpin conformation: a δ-linked ornithine turn connecting positions 17 and 36 to create a macrocycle and an intramolecular disulfide linkage between positions 24 and 29. An N-methyl group at position 33 blocks uncontrolled aggregation. The peptide readily crystallizes as a folded β-hairpin, which assembles hierarchically in the crystal lattice. Three β-hairpin monomers assemble to form a triangular trimer, four trimers assemble in a tetrahedral arrangement to form a dodecamer, and five dodecamers pack together to form an annular pore. This hierarchical assembly provides a model, in which full-length Aβ transitions from an unfolded monomer to a folded β-hairpin, which assembles to form oligomers that further pack to form an annular pore. This model may provide a better understanding of the molecular basis of Alzheimer's disease at atomic resolution.
- Department of Chemistry, University of California, Irvine , Irvine, California 92697-2025, United States.
Organizational Affiliation: