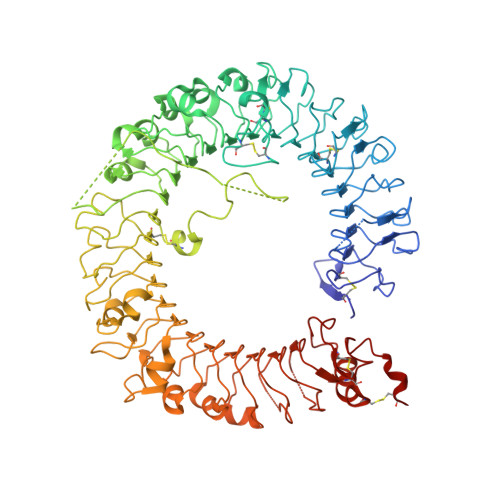
Autoinhibition and relief mechanism by the proteolytic processing of Toll-like receptor 8
Tanji, H., Ohto, U., Motoi, Y., Shibata, T., Miyake, K., Shimizu, T.(2016) Proc Natl Acad Sci U S A 113: 3012-3017
- PubMed: 26929371 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1516000113
- Primary Citation Related Structures:
5HDH - PubMed Abstract:
Toll-like receptor 8 (TLR8) senses single-stranded RNA (ssRNA) and initiates innate immune responses. TLR8 requires proteolytic cleavage at the loop region (Z-loop) between leucine-rich repeat (LRR) 14 and LRR15 for its activation. However, the molecular basis of Z-loop processing remains unknown. To elucidate the mechanism of Z-loop processing, we performed biochemical and structural studies of how the Z-loop affects the function of TLR8. TLR8 with the uncleaved Z-loop is unable to form a dimer, which is essential for activation, irrespective of the presence of agonistic ligands. Crystallographic analysis revealed that the uncleaved Z-loop located on the ascending lateral face prevents the approach of the dimerization partner by steric hindrance. This autoinhibition mechanism of dimerization by the Z-loop might be occurring in the proteins of the same subfamily, TLR7 and TLR9.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Tokyo 113-0033, Japan;
Organizational Affiliation: