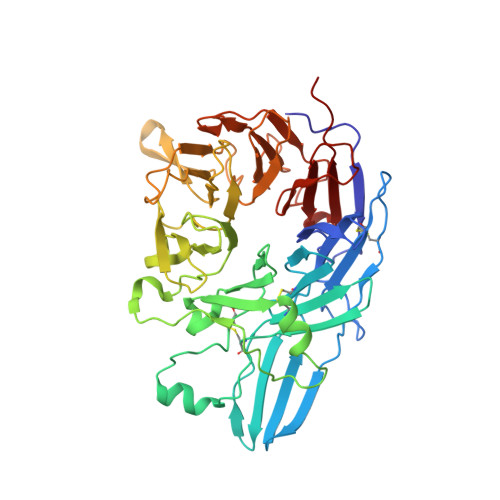
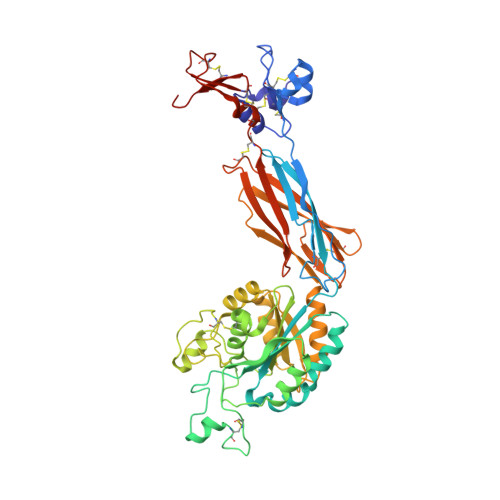
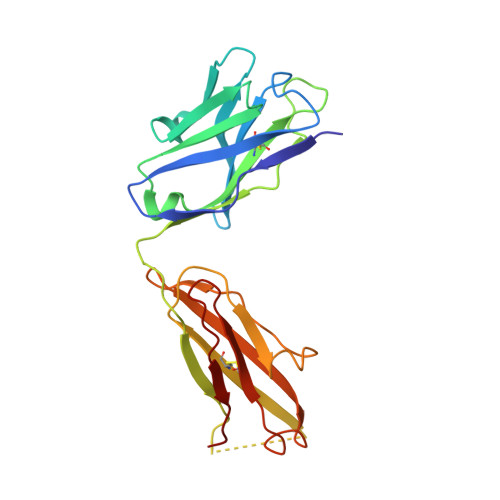
beta-Subunit Binding Is Sufficient for Ligands to Open the Integrin alpha IIb beta 3 Headpiece.
Lin, F.Y., Zhu, J., Eng, E.T., Hudson, N.E., Springer, T.A.(2016) J Biological Chem 291: 4537-4546
- PubMed: 26631735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.705624
- Primary Citation Related Structures:
4Z7N, 4Z7O, 4Z7Q, 5HDB - PubMed Abstract:
The platelet integrin αIIbβ3 binds to a KQAGDV motif at the fibrinogen γ-chain C terminus and to RGD motifs present in loops in many extracellular matrix proteins. These ligands bind in a groove between the integrin α and β-subunits; the basic Lys or Arg side chain hydrogen bonds to the αIIb-subunit, and the acidic Asp side chain coordinates to a metal ion held by the β3-subunit. Ligand binding induces headpiece opening, with conformational change in the β-subunit. During this opening, RGD slides in the ligand-binding pocket toward αIIb, with movement of the βI-domain β1-α1 loop toward αIIb, enabling formation of direct, charged hydrogen bonds between the Arg side chain and αIIb. Here we test whether ligand interactions with β3 suffice for stable ligand binding and headpiece opening. We find that the AGDV tetrapeptide from KQAGDV binds to the αIIbβ3 headpiece with affinity comparable with the RGDSP peptide from fibronectin. AGDV induced complete headpiece opening in solution as shown by increase in hydrodynamic radius. Soaking of AGDV into closed αIIbβ3 headpiece crystals induced intermediate states similarly to RGDSP. AGDV has very little contact with the α-subunit. Furthermore, as measured by epitope exposure, AGDV, like the fibrinogen γ C-terminal peptide and RGD, caused integrin extension on the cell surface. Thus, pushing by the β3-subunit on Asp is sufficient for headpiece opening and ligand sliding, and no pulling by the αIIb subunit on Arg is required.
- From the Department of Biological Chemistry and Molecular Pharmacology, Program in Cellular and Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, Massachusetts 02115.
Organizational Affiliation: