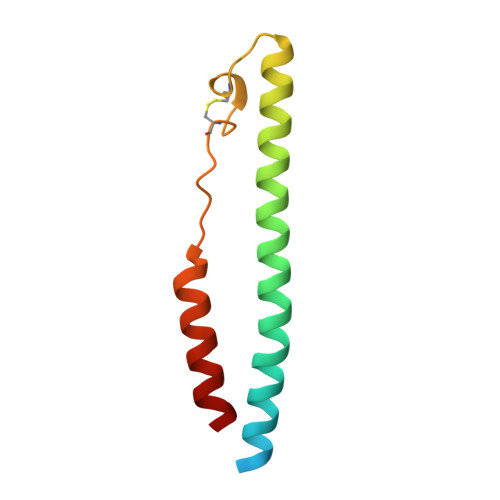
Structural characterization of a fusion glycoprotein from a retrovirus that undergoes a hybrid 2-step entry mechanism.
Aydin, H., Smrke, B.M., Lee, J.E.(2013) FASEB J 27: 5059-5071
- PubMed: 24036886 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1096/fj.13-232371
- Primary Citation Related Structures:
4JPR, 5H9C - PubMed Abstract:
Entry of enveloped viruses into host cells is mediated by their surface envelope glycoproteins (Env). On the surface of the virus, Env is in a metastable, prefusion state, primed to catalyze the fusion of the viral and host membranes. An external trigger is needed to promote the drastic conformational changes necessary for the fusion subunit to fold into the low-energy, 6-helix bundle. These triggers typically facilitate pH-independent entry at the plasma membrane or pH-dependent entry in a low-pH endosomal compartment. The α-retrovirus avian sarcoma leukosis virus (ASLV) has a rare, 2-step entry mechanism with both pH-dependent and pH-independent features. Here, we present the 2.0-Å-resolution crystal structure of the ASLV transmembrane (TM) fusion protein. Our structural and biophysical studies indicated that unlike other pH-dependent or pH-independent viral TMs, the ASLV fusion subunit is stable irrespective of pH. Two histidine residues (His490 and His492) in the chain reversal region confer stability at low pH. A structural comparison of class I viral fusion proteins suggests that the presence of a positive charge, either a histidine or arginine amino acid, stabilizes a helical dipole moment and is a signature of fusion proteins active at low pH. The structure now reveals key residues and features that explain its 2-step mechanism, and we discuss the implications of the ASLV TM structure in the context of general mechanisms required for membrane fusion.
- 11 King's College Cir., Rm. 6316, Toronto, ON, Canada M5S 1A8. jeff.lee@utoronto.ca.
Organizational Affiliation: