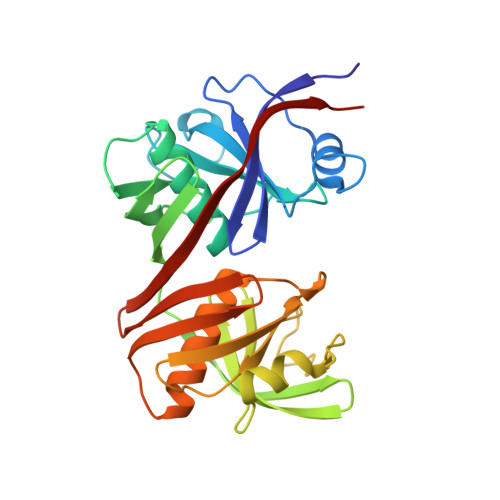
Structural basis for redox sensitivity in Corynebacterium glutamicum diaminopimelate epimerase: an enzyme involved in l-lysine biosynthesis.
Sagong, H.Y., Kim, K.J.(2017) Sci Rep 7: 42318-42318
- PubMed: 28176858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep42318
- Primary Citation Related Structures:
5H2G, 5H2Y, 5M47 - PubMed Abstract:
Diaminopimelate epimerase (DapF) is one of the crucial enzymes involved in l-lysine biosynthesis, where it converts l,l-diaminopimelate (l,l-DAP) into d,l-DAP. DapF is also considered as an attractive target for the development of antibacterial drugs. Here, we report the crystal structure of DapF from Corynebacterium glutamicum (CgDapF). Structures of CgDapF obtained under both oxidized and reduced conditions reveal that the function of CgDapF is regulated by redox-switch modulation via reversible disulfide bond formation between two catalytic cysteine residues. Under oxidized condition, two catalytic cysteine residues form a disulfide bond; these same cysteine residues exist in reduced form under reduced condition. Disulfide bond formation also induces a subsequent structural change in the dynamic catalytic loop at the active site, which results in open/closed conformational change at the active site. We also determined the crystal structure of CgDapF in complex with its product d,l-DAP, and elucidated how the enzyme recognizes its substrate l,l-DAP as a substrate. Moreover, the structure in complex with the d,l-DAP product reveals that CgDapF undergoes a large open/closed domain movement upon substrate binding, resulting in a completely buried active site with the substrate bound.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, Korea.
Organizational Affiliation: