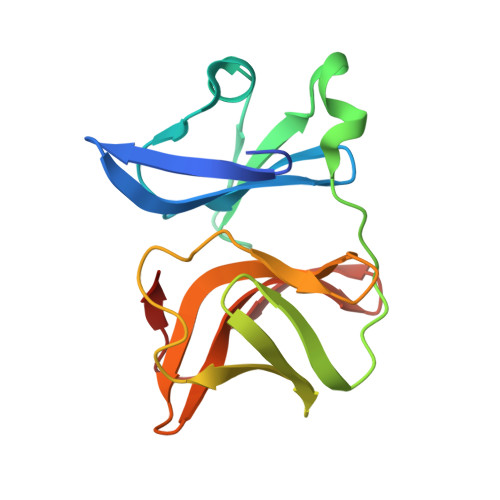
Structure-function insights into chikungunya virus capsid protein: Small molecules targeting capsid hydrophobic pocket.
Sharma, R., Kesari, P., Kumar, P., Tomar, S.(2018) Virology 515: 223-234
- PubMed: 29306785 Search on PubMed
- DOI: https://doi.org/10.1016/j.virol.2017.12.020
- Primary Citation Related Structures:
5H23 - PubMed Abstract:
The crystal structure of chikungunya (CHIKV) virus capsid protease domain has been determined at 2.2Å. Structure reveals a chymotrypsin-like protease fold with a conserved hydrophobic pocket in CHIKV capsid protein (CP) for interaction with the cytoplasmic tail of E2 (cdE2) similar to the capsid protein of other alphaviruses. Molecular contacts between CP-cdE2 were determined by fitting structures of CHIKV CP and cdE2 into the cryo-EM map of Venezuelan equine encephalitis virus (VEEV). Binding of (S)-(+)-Mandelic acid (MDA) and Ethyl 3-aminobenzoate (EAB) to the hydrophobic pocket of CP was evaluated by molecular docking. Surface plasmon resonance (SPR) and fluorescence spectroscopy experiments confirmed MDA and EAB binding to the CP. The binding constants (K D ) obtained from SPR for MDA and EAB were 1.2 × 10 -3 M and 0.2 × 10 -9 M, respectively. This study adds to the understanding of chikungunya virus structural proteins and may serve as the basis for antiviral development against chikungunya disease.
- Department of Biotechnology, Indian Institute of Technology Roorkee, Roorkee 247667, India.
Organizational Affiliation: