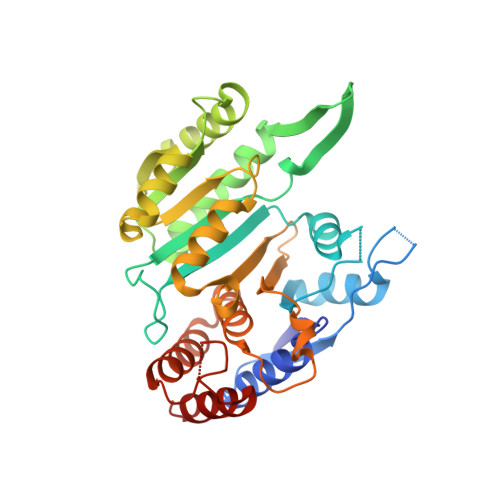
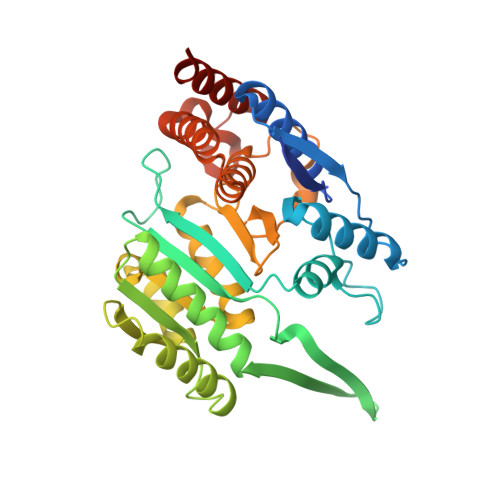
Molecular mechanism of the allosteric regulation of the alpha gamma heterodimer of human NAD-dependent isocitrate dehydrogenase.
Ma, T., Peng, Y., Huang, W., Ding, J.(2017) Sci Rep 7: 40921-40921
- PubMed: 28098230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep40921
- Primary Citation Related Structures:
5GRE, 5GRF, 5GRH, 5GRI, 5GRL - PubMed Abstract:
Human NAD-dependent isocitrate dehydrogenase catalyzes the decarboxylation of isocitrate (ICT) into α-ketoglutarate in the Krebs cycle. It exists as the α 2 βγ heterotetramer composed of the αβ and αγ heterodimers. Previously, we have demonstrated biochemically that the α 2 βγ heterotetramer and αγ heterodimer can be allosterically activated by citrate (CIT) and ADP. In this work, we report the crystal structures of the αγ heterodimer with the γ subunit bound without or with different activators. Structural analyses show that CIT, ADP and Mg 2+ bind adjacent to each other at the allosteric site. The CIT binding induces conformational changes at the allosteric site, which are transmitted to the active site through the heterodimer interface, leading to stabilization of the ICT binding at the active site and thus activation of the enzyme. The ADP binding induces no further conformational changes but enhances the CIT binding through Mg 2+ -mediated interactions, yielding a synergistic activation effect. ICT can also bind to the CIT-binding subsite, which induces similar conformational changes but exhibits a weaker activation effect. The functional roles of the key residues are verified by mutagenesis, kinetic and structural studies. Our structural and functional data together reveal the molecular mechanism of the allosteric regulation of the αγ heterodimer.
- National Center for Protein Science Shanghai, State Key Laboratory of Molecular Biology, Center for Excellence in Molecular Cell Science, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yueyang Road, Shanghai 200031, China.
Organizational Affiliation: