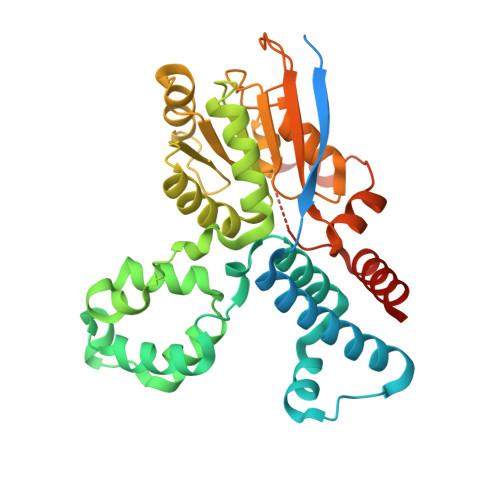
Crystal structure of the ATPase-C domain of the chromatin remodeller Fun30 from Saccharomyces cerevisiae.
Liu, L., Jiang, T.(2017) Acta Crystallogr F Struct Biol Commun 73: 9-15
- PubMed: 28045388 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16019269
- Primary Citation Related Structures:
5GN1 - PubMed Abstract:
Fun30 (Function unknown now 30) is a chromatin remodeller belonging to the Snf2 family. It has previously been reported to be a regulator of several cellular activities, including DNA repair, gene silencing and maintenance of chromatin structure. Here, the crystal structure of the Fun30 ATPase-C domain (the C-lobe of the ATPase domain) is reported at 1.95 Å resolution. Although the structure displays overall similarities to those of other Snf2 family members, a new structural module was found to be specific to the Fun30 subfamily. Fun30 ATPase-C was shown be monomeric in solution and showed no detectable affinity for dsDNA.
- National Laboratory of Biomacromolecules, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Datun Road, Chaoyang District, Beijing 100101, People's Republic of China.
Organizational Affiliation: