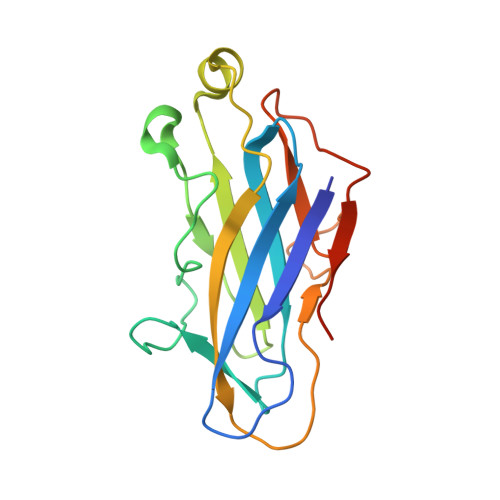
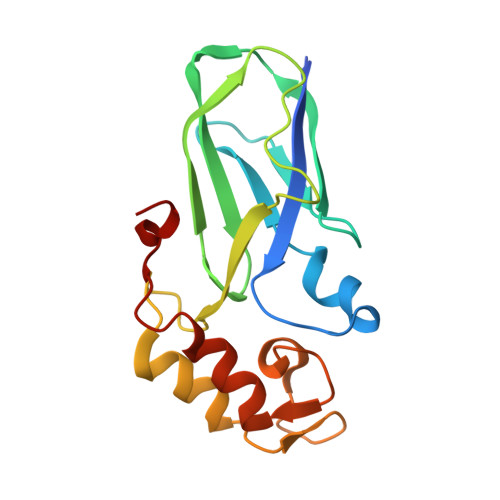
Diverse specificity of cellulosome attachment to the bacterial cell surface.
Bras, J.L., Pinheiro, B.A., Cameron, K., Cuskin, F., Viegas, A., Najmudin, S., Bule, P., Pires, V.M., Romao, M.J., Bayer, E.A., Spencer, H.L., Smith, S., Gilbert, H.J., Alves, V.D., Carvalho, A.L., Fontes, C.M.(2016) Sci Rep 6: 38292-38292
- PubMed: 27924829 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep38292
- Primary Citation Related Structures:
5G5D, 5K39, 5M0Y - PubMed Abstract:
During the course of evolution, the cellulosome, one of Nature's most intricate multi-enzyme complexes, has been continuously fine-tuned to efficiently deconstruct recalcitrant carbohydrates. To facilitate the uptake of released sugars, anaerobic bacteria use highly ordered protein-protein interactions to recruit these nanomachines to the cell surface. Dockerin modules located within a non-catalytic macromolecular scaffold, whose primary role is to assemble cellulosomal enzymatic subunits, bind cohesin modules of cell envelope proteins, thereby anchoring the cellulosome onto the bacterial cell. Here we have elucidated the unique molecular mechanisms used by anaerobic bacteria for cellulosome cellular attachment. The structure and biochemical analysis of five cohesin-dockerin complexes revealed that cell surface dockerins contain two cohesin-binding interfaces, which can present different or identical specificities. In contrast to the current static model, we propose that dockerins utilize multivalent modes of cohesin recognition to recruit cellulosomes to the cell surface, a mechanism that maximises substrate access while facilitating complex assembly.
- Centro Interdisciplinar de Investigação em Sanidade Animal, Faculdade de Medicina Veterinária, Universidade de Lisboa, 1300-477 Lisboa, Portugal.
Organizational Affiliation: