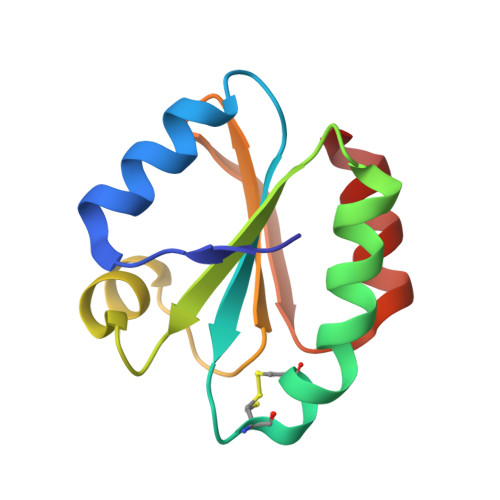
Is dimerization a common feature in thioredoxins? The case of thioredoxin from Litopenaeus vannamei.
Campos-Acevedo, A.A., Sotelo-Mundo, R.R., Perez, J., Rudino-Pinera, E.(2017) Acta Crystallogr D Struct Biol 73: 326-339
- PubMed: 28375144 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798317002066
- PubMed Abstract:
The quaternary structure of the redox protein thioredoxin (Trx) has been debated. For bacterial Trx, there is no question regarding its monomeric state. In humans and other eukaryotes, the presence of a cysteine residue at the crystallographic symmetry axis points to the relevance of dimer formation in solution and in vivo. Crystallographic data for shrimp thioredoxin (LvTrx) obtained under different redox conditions reveal a dimeric arrangement mediated by a disulfide bond through residue Cys73 and other hydrophobic interactions located in the crystallographic interface, as reported for human Trx. Through the analysis of five mutants located at the crystallographic interface, this study provides structural and biochemical evidence for the existence in solution of monomeric and dimeric populations of wild-type LvTrx and five mutants. Based on the results of biochemical assays, SAXS studies and the crystallographic structures of three of the studied mutants (Cys73Ser, Asp60Ser and Trp31Ala), it is clear that the Cys73 residue is essential for dimerization. However, its mutation to Ser produces an enzyme which has similar redox activity in vitro to the wild type. A putative regulatory function of dimerization is proposed based on structural analysis. Nonetheless, the biological role of LvTrx dimerization needs to be experimentally unveiled. Additionally, the findings of this work reopen the discussion regarding the existence of similar behaviour in human thioredoxin, which shares a Cys at position 73 with LvTrx, a structural feature that is also present in some Trxs from vertebrates and crustaceans.
- Departamento de Medicina Molecular y Bioprocesos, Instituto de Biotecnología (IBT), Universidad Nacional Autónoma de México (UNAM), Avenida Universidad 2001, Colonia Chamilpa, 62210 Cuernavaca, MOR, Mexico.
Organizational Affiliation: