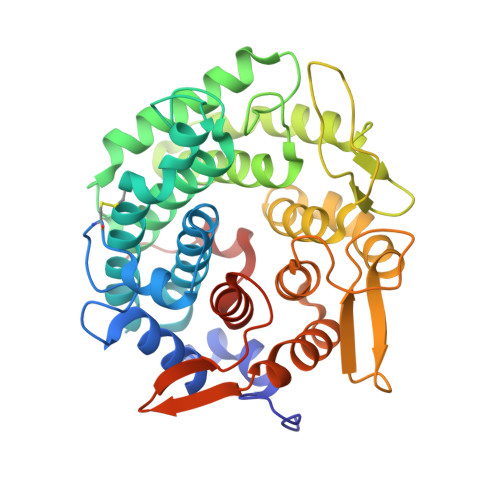
Structural and functional insights into asymmetric enzymatic dehydration of alkenols.
Nestl, B.M., Geinitz, C., Popa, S., Rizek, S., Haselbeck, R.J., Stephen, R., Noble, M.A., Fischer, M.P., Ralph, E.C., Hau, H.T., Man, H., Omar, M., Turkenburg, J.P., van Dien, S., Culler, S.J., Grogan, G., Hauer, B.(2017) Nat Chem Biol 13: 275-281
- PubMed: 28068311 Search on PubMed
- DOI: https://doi.org/10.1038/nchembio.2271
- Primary Citation Related Structures:
5G1U, 5G1V, 5G1W - PubMed Abstract:
The asymmetric dehydration of alcohols is an important process for the direct synthesis of alkenes. We report the structure and substrate specificity of the bifunctional linalool dehydratase isomerase (LinD) from the bacterium Castellaniella defragrans that catalyzes in nature the hydration of β-myrcene to linalool and the subsequent isomerization to geraniol. Enzymatic kinetic resolutions of truncated and elongated aromatic and aliphatic tertiary alcohols (C5-C15) that contain a specific signature motif demonstrate the broad substrate specificity of LinD. The three-dimensional structure of LinD from Castellaniella defragrans revealed a pentamer with active sites at the protomer interfaces. Furthermore, the structure of LinD in complex with the product geraniol provides initial mechanistic insights into this bifunctional enzyme. Site-directed mutagenesis confirmed active site amino acid residues essential for its dehydration and isomerization activity. These structural and mechanistic insights facilitate the development of hydrating catalysts, enriching the toolbox for novel bond-forming biocatalysis.
- Institute of Technical Biochemistry, Universitaet Stuttgart, Stuttgart, Germany.
Organizational Affiliation: