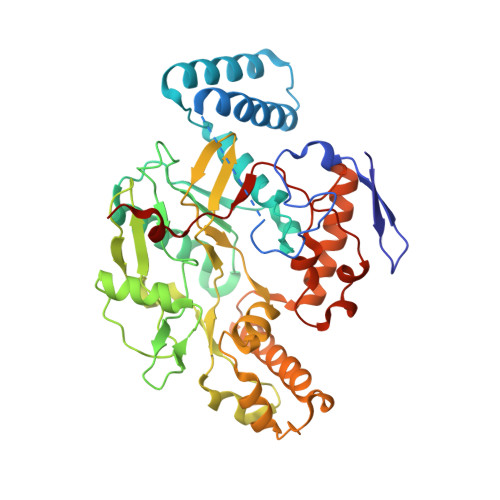
Electrostatic Control of Isoform Selective Inhibitor Binding in Nitric Oxide Synthase.
Li, H., Wang, H., Kang, S., Silverman, R.B., Poulos, T.L.(2016) Biochemistry 55: 3702
- PubMed: 27250740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00261
- Primary Citation Related Structures:
5G0N, 5G0O, 5G0P - PubMed Abstract:
Development of potent and isoform selective nitric oxide synthase (NOS) inhibitors is challenging because of the structural similarity in the heme active sites. One amino acid difference between NOS isoforms, Asp597 in rat neuronal NOS (nNOS) versus Asn368 in bovine endothelial NOS (eNOS), has been identified as the structural basis for why some dipeptide amide inhibitors bind more tightly to nNOS than to eNOS. We now have found that the same amino acid variation is responsible for substantially different binding modes and affinity for a new class of aminopyridine-based inhibitors.
- Departments of Molecular Biology and Biochemistry, Pharmaceutical Sciences, and Chemistry, University of California , Irvine, California 92697-3900, United States.
Organizational Affiliation: