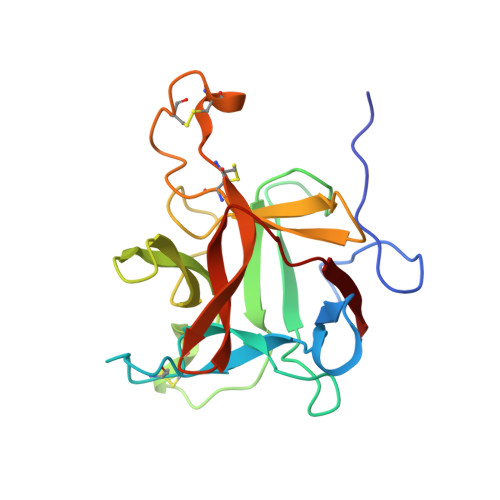
Structures of a bi-functional Kunitz-type STI family inhibitor of serine and aspartic proteases: Could the aspartic protease inhibition have evolved from a canonical serine protease-binding loop?
Guerra, Y., Valiente, P.A., Pons, T., Berry, C., Rudino-Pinera, E.(2016) J Struct Biol 195: 259-271
- PubMed: 27329566 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2016.06.014
- Primary Citation Related Structures:
5FNW, 5FNX, 5FZU, 5FZY, 5FZZ, 5G00 - PubMed Abstract:
Bi-functional inhibitors from the Kunitz-type soybean trypsin inhibitor (STI) family are glycosylated proteins able to inhibit serine and aspartic proteases. Here we report six crystal structures of the wild-type and a non-glycosylated mutant of the bifunctional inhibitor E3Ad obtained at different pH values and space groups. The crystal structures show that E3Ad adopts the typical β-trefoil fold of the STI family exhibiting some conformational changes due to pH variations and crystal packing. Despite the high sequence identity with a recently reported potato cathepsin D inhibitor (PDI), three-dimensional structures obtained in this work show a significant conformational change in the protease-binding loop proposed for aspartic protease inhibition. The E3Ad binding loop for serine protease inhibition is also proposed, based on structural similarity with a novel non-canonical conformation described for the double-headed inhibitor API-A from the Kunitz-type STI family. In addition, structural and sequence analyses suggest that bifunctional inhibitors of serine and aspartic proteases from the Kunitz-type STI family are more similar to double-headed inhibitor API-A than other inhibitors with a canonical protease-binding loop.
- Departamento de Medicina Molecular y Bioprocesos, Instituto de Biotecnología, Universidad Nacional Autónoma de México (UNAM), Avenida Universidad 2001, Colonia Chamilpa, Cuernavaca, Morelos CP 62210, Mexico. Electronic address: yaselg@ibt.unam.mx.
Organizational Affiliation: