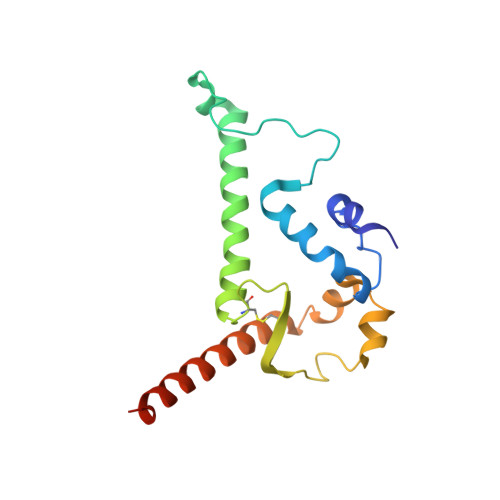
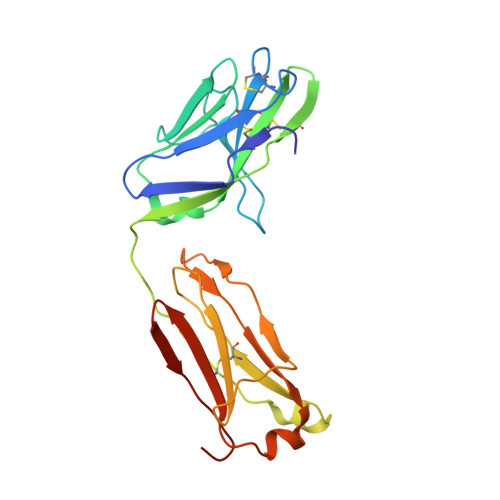
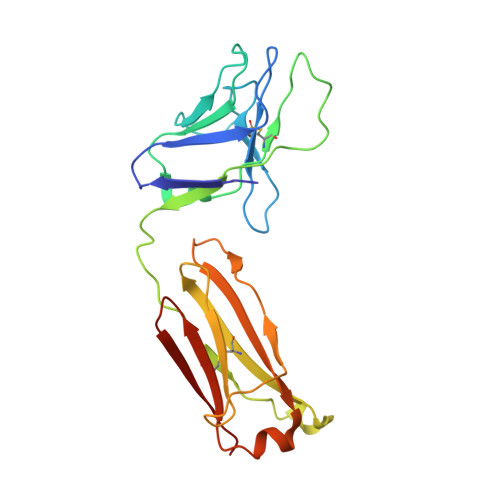
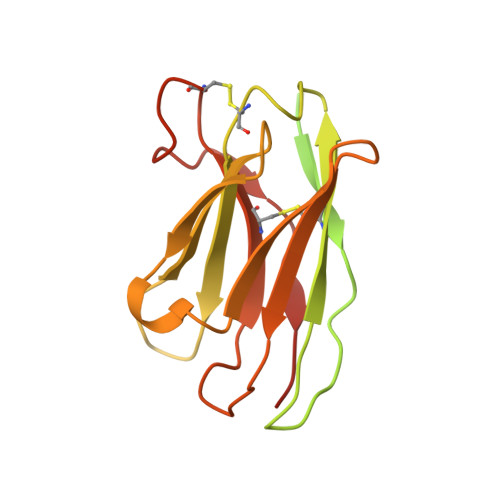
Trimeric HIV-1-Env Structures Define Glycan Shields from Clades A, B and G
Stewart-Jones, G.B.E., Soto, C., Lemmin, T., Chuang, G.Y., Druz, A., Kong, R., Thomas, P.V., Wagh, K., Zhou, T., Behrens, A.J., Bylund, T., Choi, C.W., Davison, J.R., Georgiev, I.S., Joyce, M.G., Kwon, Y.D., Pancera, M., Taft, J., Yang, Y., Zhang, B., Shivatare, S.S., Shivatare, V.S., Lee, C.C.D., Wu, C.Y., Bewley, C.A., Burton, D.R., Koff, W.C., Connors, M., Crispin, M., Korber, B.T., Wong, C.H., Mascola, J.R., Kwong, P.D.(2016) Cell 165: 813
- PubMed: 27114034 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2016.04.010
- Primary Citation Related Structures:
5FYJ, 5FYK, 5FYL - PubMed Abstract:
The HIV-1-envelope (Env) trimer is covered by a glycan shield of ∼90 N-linked oligosaccharides, which comprises roughly half its mass and is a key component of HIV evasion from humoral immunity. To understand how antibodies can overcome the barriers imposed by the glycan shield, we crystallized fully glycosylated Env trimers from clades A, B, and G, visualizing the shield at 3.4-3.7 Å resolution. These structures reveal the HIV-1-glycan shield to comprise a network of interlocking oligosaccharides, substantially ordered by glycan crowding, that encase the protein component of Env and enable HIV-1 to avoid most antibody-mediated neutralization. The revealed features delineate a taxonomy of N-linked glycan-glycan interactions. Crowded and dispersed glycans are differently ordered, conserved, processed, and recognized by antibody. The structures, along with glycan-array binding and molecular dynamics, reveal a diversity in oligosaccharide affinity and a requirement for accommodating glycans among known broadly neutralizing antibodies that target the glycan-shielded trimer.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: