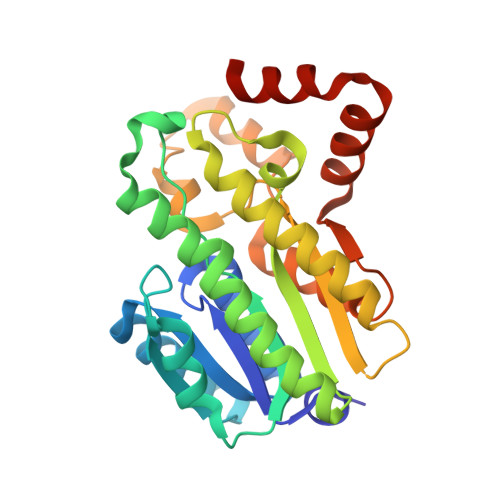
Structural and Biochemical Insights Into 7Beta-Hydroxysteroid Dehydrogenase Stereoselectivity.
Savino, S., Ferrandi, E.E., Forneris, F., Rovida, S., Riva, S., Monti, D., Mattevi, A.(2016) Proteins 84: 859
- PubMed: 27006087 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25036
- Primary Citation Related Structures:
5FYD - PubMed Abstract:
Hydroxysteroid dehydrogenases are of great interest as biocatalysts for transformations involving steroid substrates. They feature a high degree of stereo- and regio-selectivity, acting on a defined atom with a specific configuration of the steroid nucleus. The crystal structure of 7β-hydroxysteroid dehydrogenase from Collinsella aerofaciens reveals a loop gating active-site accessibility, the bases of the specificity for NADP(+) , and the general architecture of the steroid binding site. Comparison with 7α-hydroxysteroid dehydrogenase provides a rationale for the opposite stereoselectivity. The presence of a C-terminal extension reshapes the substrate site of the β-selective enzyme, possibly leading to an inverted orientation of the bound substrate. Proteins 2016; 84:859-865. © 2016 Wiley Periodicals, Inc.
- Department of Biology and Biotechnology, University of Pavia, via Ferrata 9, Pavia, 27100, Italy.
Organizational Affiliation: